The complete INTEGRATE workflow: from data to posterior¶
[1]:
try:
# Check if the code is running in an IPython kernel (which includes Jupyter notebooks)
get_ipython()
# If the above line doesn't raise an error, it means we are in a Jupyter environment
# Execute the magic commands using IPython's run_line_magic function
get_ipython().run_line_magic('load_ext', 'autoreload')
get_ipython().run_line_magic('autoreload', '2')
except:
# If get_ipython() raises an error, we are not in a Jupyter environment
# # # # # #%load_ext autoreload
# # # # # #%autoreload 2
pass
[2]:
import os
import integrate as ig
from geoprior1d import geoprior1d
hardcopy = True
import matplotlib.pyplot as plt
import numpy as np
import copy
[3]:
N = 100_000 # Number of prior model realizations to generate (this is just for testing, use a larger number for better results)
GETTING THE DATA AND GEX FILE for gthe chosen area¶
[4]:
case = 'DAUGAARD'
f_xlsx_files = ['daugaard_standard.xlsx','daugaard_valley.xlsx']
ig.get_case_data(case=case, filelist=['daugaard_standard.xlsx','daugaard_valley.xlsx'])
files = ig.get_case_data(case=case)
f_data_h5 = files[0]
file_gex= ig.get_gex_file_from_data(f_data_h5)
loadFromXyz=True
if loadFromXyz:
ig.get_case_data(case=case, loadType='xyz')
useRaw = False
if useRaw:
# Unprocessed and unstacked data
file_xyz_list=['tTEM_20230727_RAW_export.xyz', 'tTEM_20230814_RAW_export.xyz', 'tTEM_20230829_RAW_export.xyz', 'tTEM_20230913_RAW_export.xyz', 'tTEM_20231109_RAW_export.xyz']
else:
# Processed and stacked data
file_xyz_list=['tTEM_20230727_AVG_export.xyz', 'tTEM_20230814_AVG_export.xyz', 'tTEM_20230829_AVG_export.xyz', 'tTEM_20230913_AVG_export.xyz', 'tTEM_20231109_AVG_export.xyz']
f_data_h5 = ig.xyz_to_h5(file_xyz_list, file_gex, showInfo=1)
print("Using data file: %s" % f_data_h5)
print("Using GEX file: %s" % file_gex)
f_data_old_h5 = f_data_h5
f_data_h5 = f_data_h5.replace('.h5', '_WF.h5')
ig.copy_hdf5_file(f_data_old_h5, f_data_h5)
Getting data for case: DAUGAARD
--> Got data for case: DAUGAARD
Getting data for case: DAUGAARD
--> Got data for case: DAUGAARD
Getting data for case: DAUGAARD
--> Got data for case: DAUGAARD
Converting XYZ to HDF5: ['tTEM_20230727_AVG_export.xyz', 'tTEM_20230814_AVG_export.xyz', 'tTEM_20230829_AVG_export.xyz', 'tTEM_20230913_AVG_export.xyz', 'tTEM_20231109_AVG_export.xyz'] → (derived from XYZ)
header [General] parsed
header [Channel1] parsed
header [Channel2] parsed
LM gates used: 14 (gates 2-15)
HM gates used: 26 (gates 5-30)
Reading tTEM_20230727_AVG_export.xyz
Reading tTEM_20230814_AVG_export.xyz
Reading tTEM_20230829_AVG_export.xyz
Reading tTEM_20230913_AVG_export.xyz
Reading tTEM_20231109_AVG_export.xyz
Removed 6 all-NaN soundings (11687 remaining)
Data has 11687 stations and 40 channels
Checking INTEGRATE data in tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export.h5
Creating UTMX
Creating UTMY
Creating LINE
Creating ELEVATION
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export.h5:D1
Using data file: tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export.h5
Using GEX file: TX07_20231016_2x4_RC20-33.gex
[4]:
'tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF.h5'
the PRIOR model(s) :¶
simulate prior model realizations using geoprior1d # https://github.com/GEUSjesper/geoprior1d We are using two prior models representing expected variability outstide and inside a buried valley system.
[5]:
f_prior_h5_list = []
for file_xlsx in f_xlsx_files:
# get filename without extension for naming the output hdf5 file
fname = file_xlsx.split('.')[0]
f_prior_h5, flags = geoprior1d(file_xlsx, Nreals=N, dz=1, dmax =90, output_file='%s_prior_N%d.h5' % (fname, N))
f_prior_h5_list.append(f_prior_h5)
# Merge the two prior hdf5 files into one, which will be used for the rest of the workflow. The merged file will be named 'daugaard_merged_prior_N10000.h5' (or with the appropriate N value).
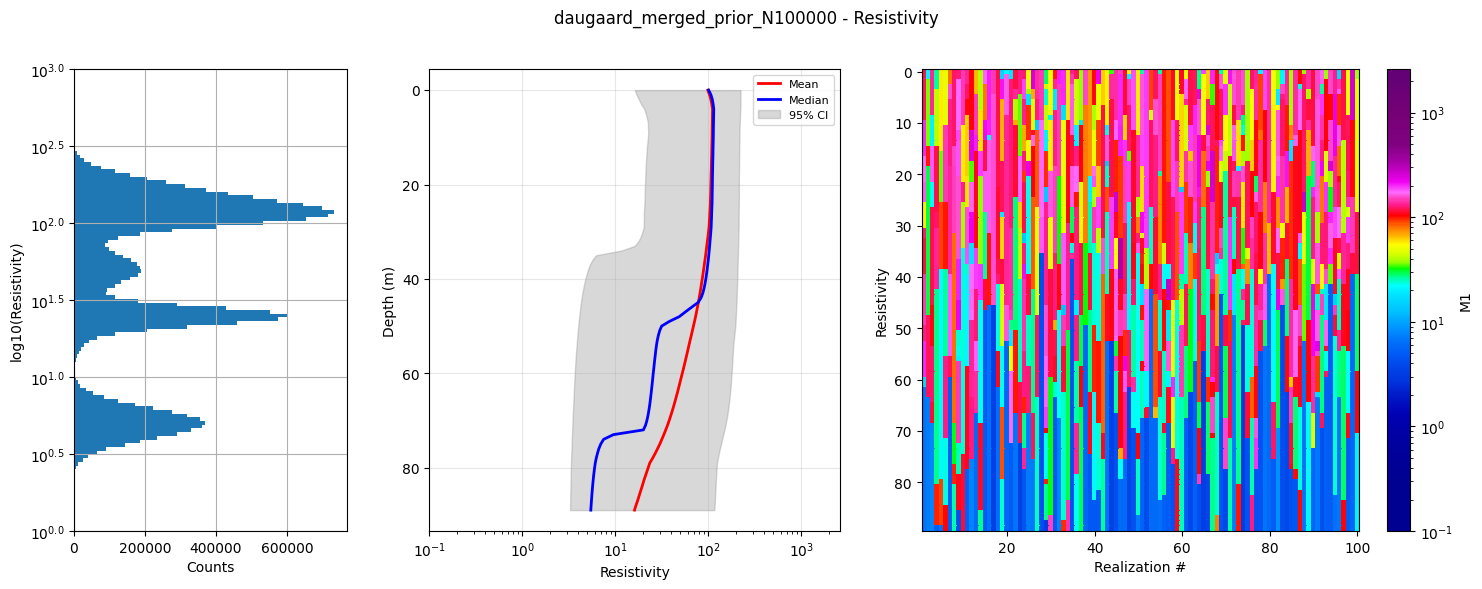
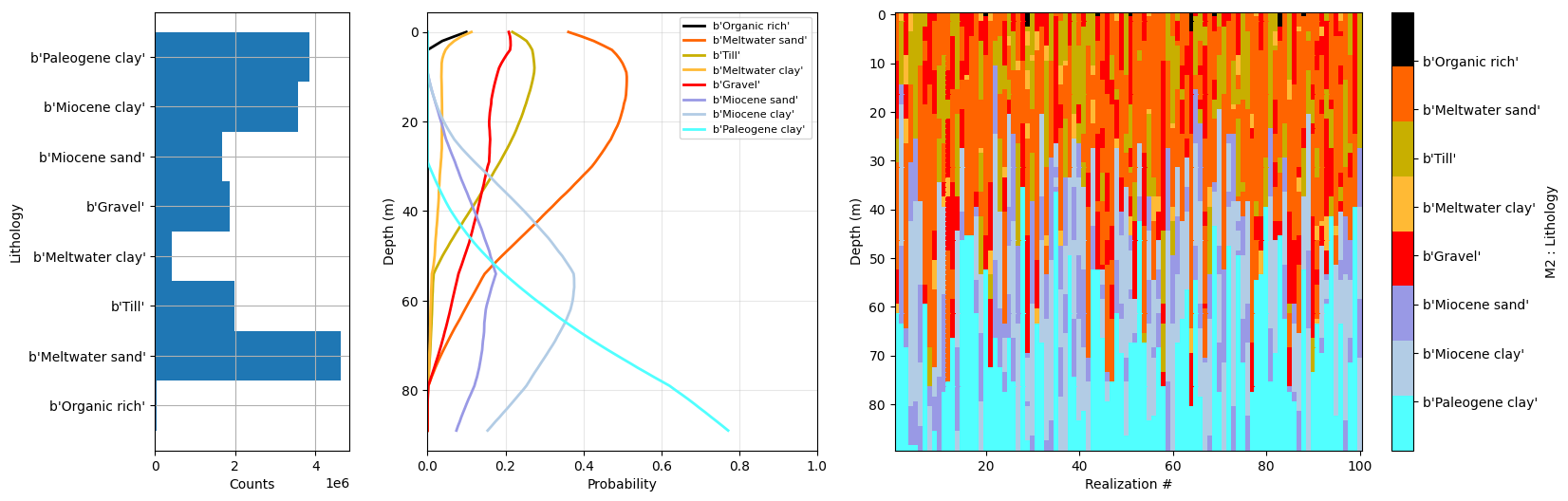
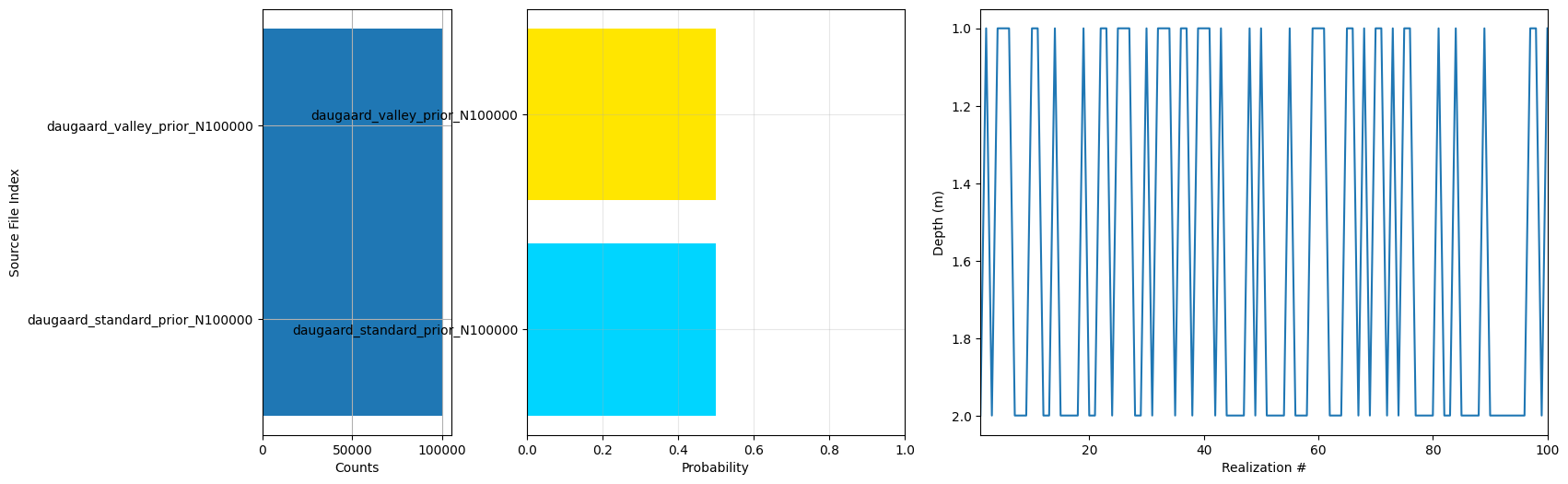
f_prior_h5 = ig.merge_prior(f_prior_h5_list, f_prior_merged_h5='daugaard_merged_prior_N%d.h5' % N)
ig.plot_prior_stats(f_prior_h5, hardcopy=hardcopy)
Using 32 parallel processes...
Generating priors: 100%|███████████████████████████████████████████████████████████████████████████| 100000/100000 [00:05<00:00, 19451.37real/s]
Prior generation completed in 5 seconds.
Using 32 parallel processes...
Generating priors: 100%|███████████████████████████████████████████████████████████████████████████| 100000/100000 [00:05<00:00, 19513.73real/s]
Prior generation completed in 5 seconds.
the tTEM DATA¶
[6]:
useLogData = False # Whether to transform data to log10 space (recommended for resistivity data)
useCorrleatedNoise = False # Whether to use correlated noise (instead of uncorrelated) when generating the prior data. This can be more realistic for geophysical data, but it also increases the computational cost.
inflateNoise = 2 # Factor to increase noise level (std) in the data, to
[7]:
if useLogData:
f_data_h5_org = f_data_h5
fname_data = f_data_h5.split('.')[0]
f_data_h5 = '%s_LOGSPACE.h5' % fname_data
ig.copy_hdf5_file(f_data_h5_org, f_data_h5)
DATA = ig.load_data(f_data_h5_org)
D_obs = DATA['d_obs'][0]
D_std = DATA['d_std'][0]
lD_obs = np.log10(D_obs)
lD_std_up = np.abs(np.log10(D_obs+D_std)-lD_obs)
lD_std_down = np.abs(np.log10(D_obs-D_std)-lD_obs)
corr_std = 0.02
lD_std = np.abs((lD_std_up+lD_std_down)/2) + corr_std
if useCorrleatedNoise:
# MISSING
pass
else:
ig.save_data_gaussian(lD_obs, D_std = lD_std, f_data_h5 = f_data_h5, id=1, showInfo=0, is_log=1)
[8]:
if inflateNoise != 1:
gf=inflateNoise
print("="*60)
print("Increasing noise level (std) by a factor of %d" % gf)
print("="*60)
D = ig.load_data(f_data_h5)
D_obs = D['d_obs'][0]
D_std = D['d_std'][0]*gf
f_data_old_h5 = f_data_h5
fname_data = f_data_h5.split('.')[0]
f_data_h5 = '%s_gf%g.h5' % (fname_data, gf)
ig.copy_hdf5_file(f_data_old_h5, f_data_h5)
ig.save_data_gaussian(D_obs, D_std=D_std, f_data_h5=f_data_h5, file_gex=file_gex)
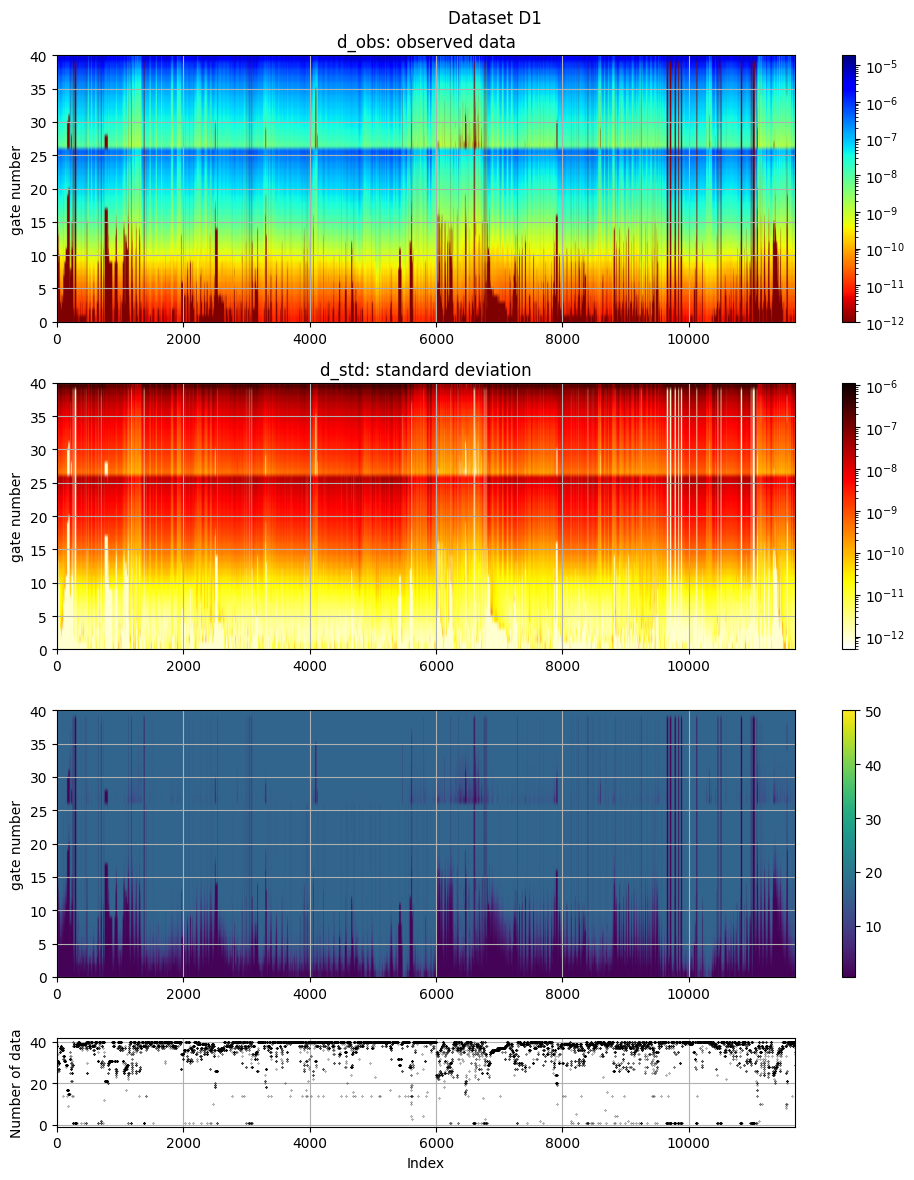
# The electromagnetic data (d_obs and d_std) can be plotted using ig.plot_data:
ig.plot_data(f_data_h5, hardcopy=hardcopy)
# Plot data channel 15 in an XY grid
ig.plot_data_xy(f_data_h5, data_channel=15, cmap='jet');
============================================================
Increasing noise level (std) by a factor of 2
============================================================
Loading data from tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF.h5. Using data types: [1]
- D1: id_prior=1, gaussian, Using 11687/40 data
Data has 11687 stations and 40 channels
Removing group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D1
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D1
plot_data: Found data set D1
plot_data: Using data set D1
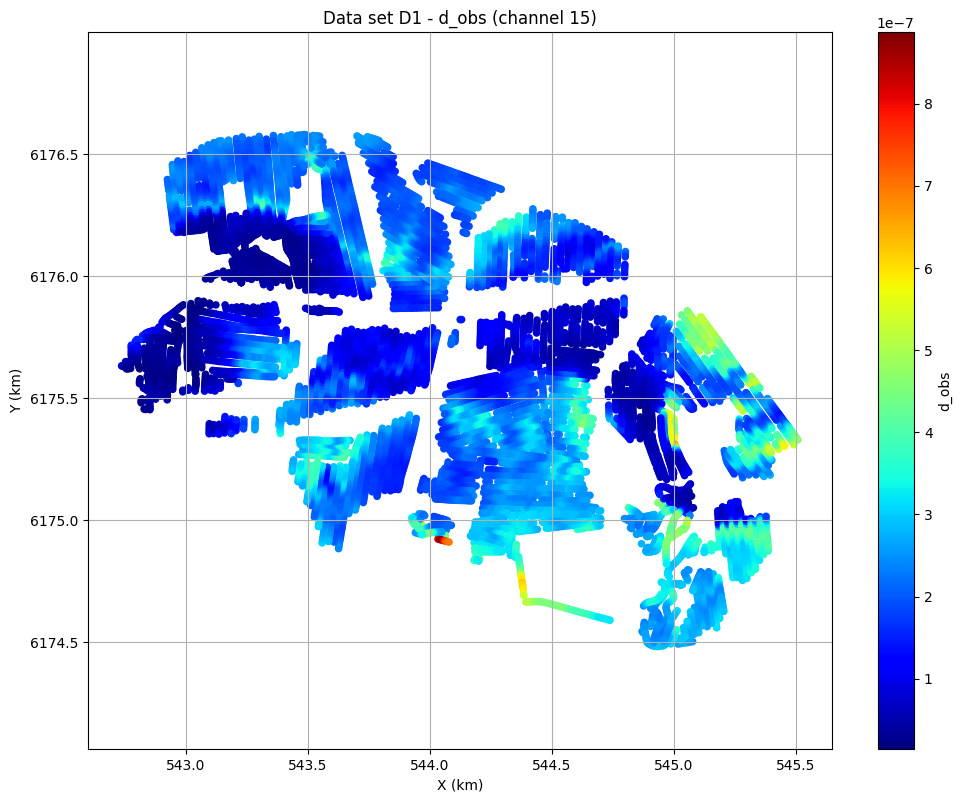
the tTEM DATA¶
[9]:
doLoadBoreholes = True
if doLoadBoreholes:
BHOLES = ig.read_borehole('daugaard_12boreholes.json', showInfo=1)
else:
BHOLES=[]
P_single=0.9 # Probability assigned to the observed class in the prior model (for discrete parameters)
BH = {}
BH['depth_top'] = [0 , 13.2, 16.6]
BH['depth_bottom'] = [12.2, 15.6, 20]
BH['class_obs'] = [2, 5, 3]
BH['class_prob'] = [P_single, P_single, P_single]
BH['X'] = 542983.01
BH['Y'] = 6175822.76
BH['name'] = 'DAU02 - Compressed'
BHOLES.append(BH.copy())
BH = {}
BH = BH.copy()
BH['depth_top'] = [0, 8, 15, 17, 23.5, 45]
BH['depth_bottom'] = [7, 14, 16, 22.5, 44, 46]
BH['class_obs'] = [3, 5, 3, 5, 6, 7]
BH['class_prob'] = [P_single, P_single, P_single, .99, P_single, P_single]
BH['X'] = 543584.098
BH['Y'] = 6175788.478
BH['name'] = '116.1602 - Compressed'
BHOLES.append(BH.copy())
# Write all boreholes together in one JSON file
ig.write_borehole(BHOLES, 'daugaard_boreholes.json', showInfo=1)
for BH in BHOLES:
print(f" {BH['name']:30s} {len(BH['depth_top'])} intervals X={BH['X']:.1f} Y={BH['Y']:.1f}")
read_borehole: read 12 borehole(s) from daugaard_12boreholes.json
DAU01 5 intervals X=544584.4 Y=6176275.2
DAU02 6 intervals X=542983.0 Y=6175822.8
DAU03 8 intervals X=544807.0 Y=6175597.0
DAU04 10 intervals X=543478.6 Y=6175871.2
DAU05 4 intervals X=544599.0 Y=6175715.0
DAU06 3 intervals X=543321.0 Y=6176073.0
DAU07 5 intervals X=543929.0 Y=6175551.0
116. 1602 7 intervals X=543584.1 Y=6175788.5
116. 535 3 intervals X=544041.1 Y=6175565.5
116. 1866 12 intervals X=543941.9 Y=6175709.7
116. 1867 4 intervals X=543712.2 Y=6175597.7
116. 1868 2 intervals X=543290.3 Y=6175528.5
[10]:
# Plot without prior info – classes labelled by numeric ID, default colours
ig.plot_boreholes(BHOLES)
# Plot without prior info
ig.plot_boreholes(BHOLES, f_prior_h5)
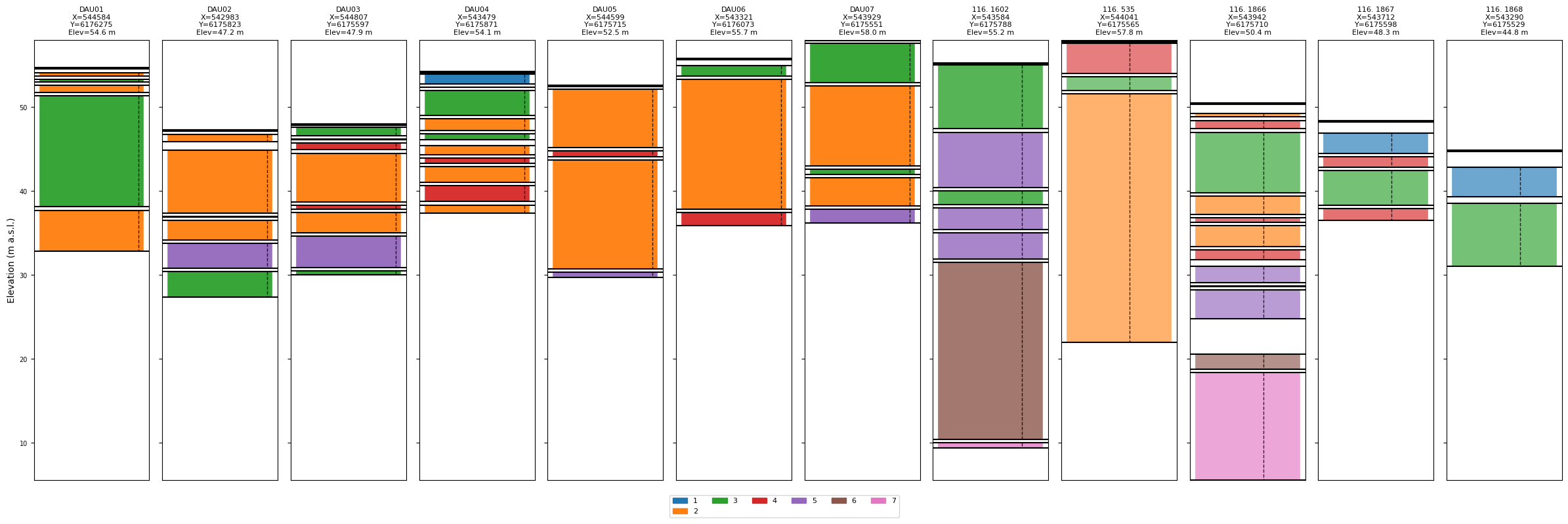
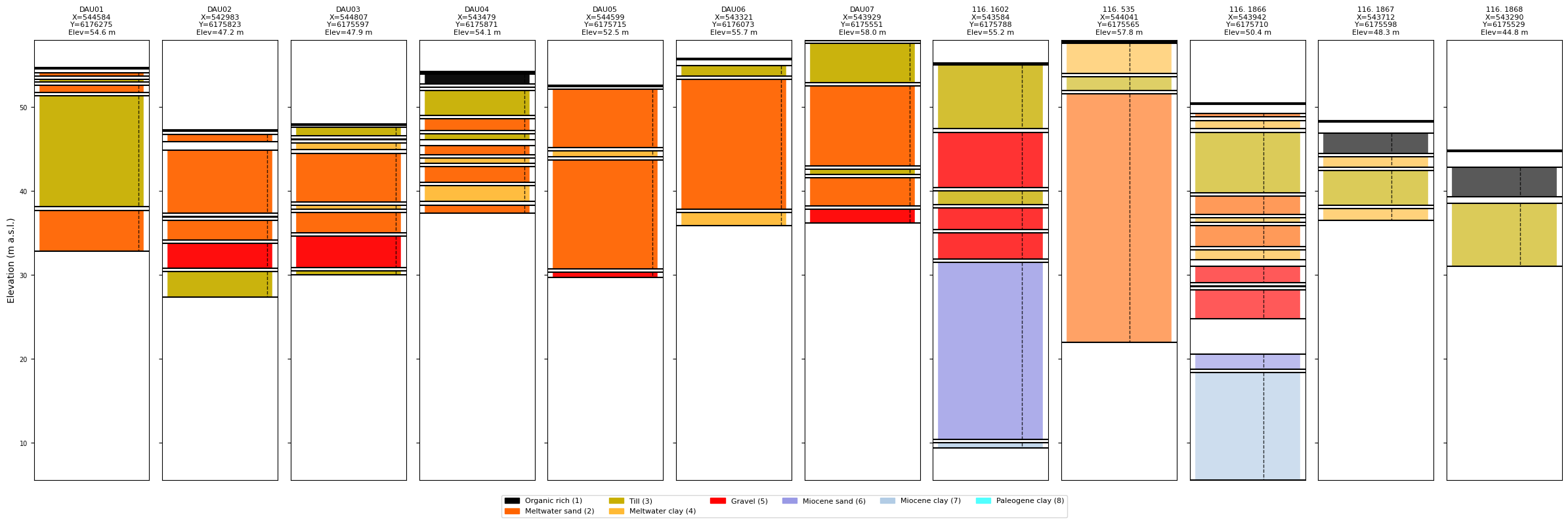
[10]:
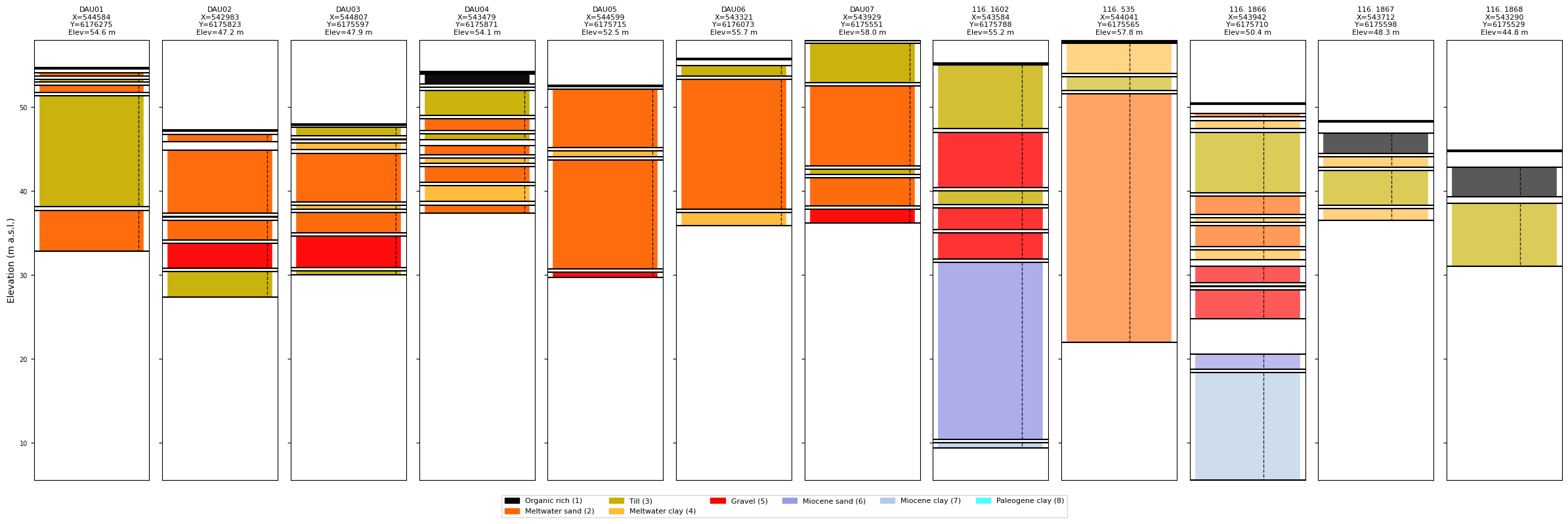
COMPUTE PRIOR DATA !!¶
[11]:
f_prior_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex, doMakePriorCopy=False)
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time=427.3s/200000 soundings. 2.1ms/sounding, 468.1it/s
[ ]:
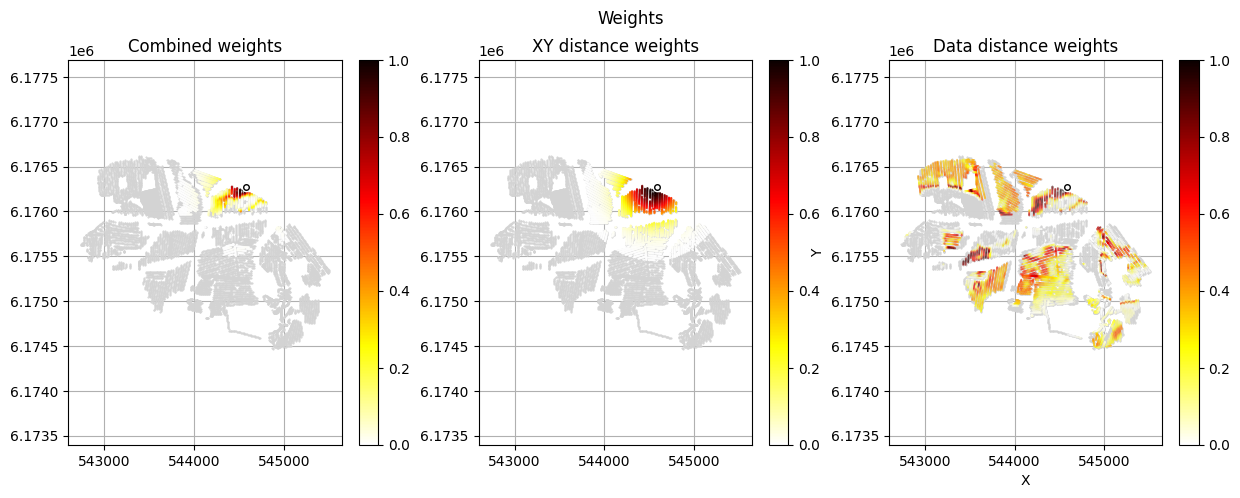
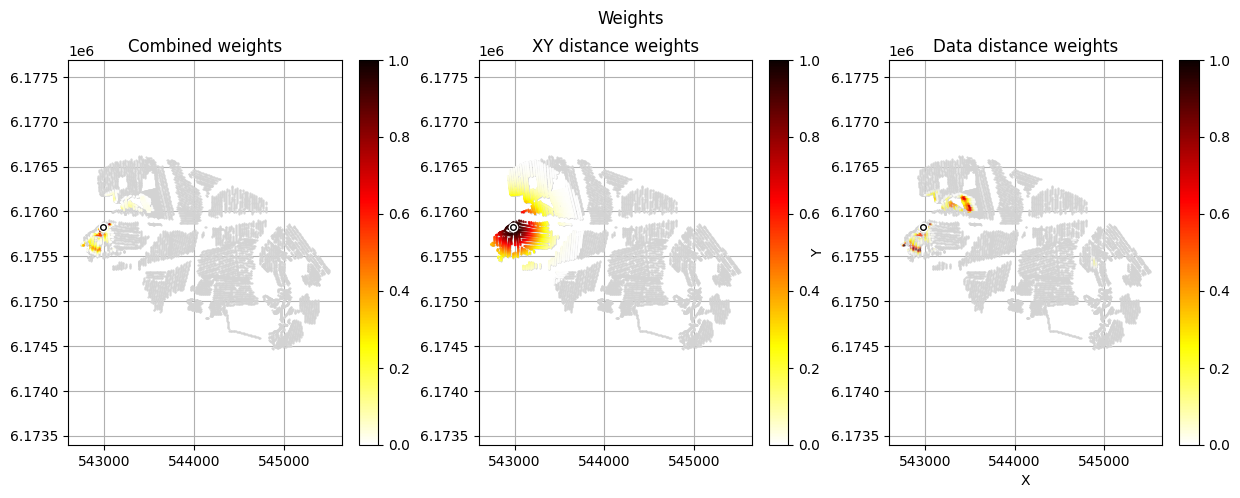
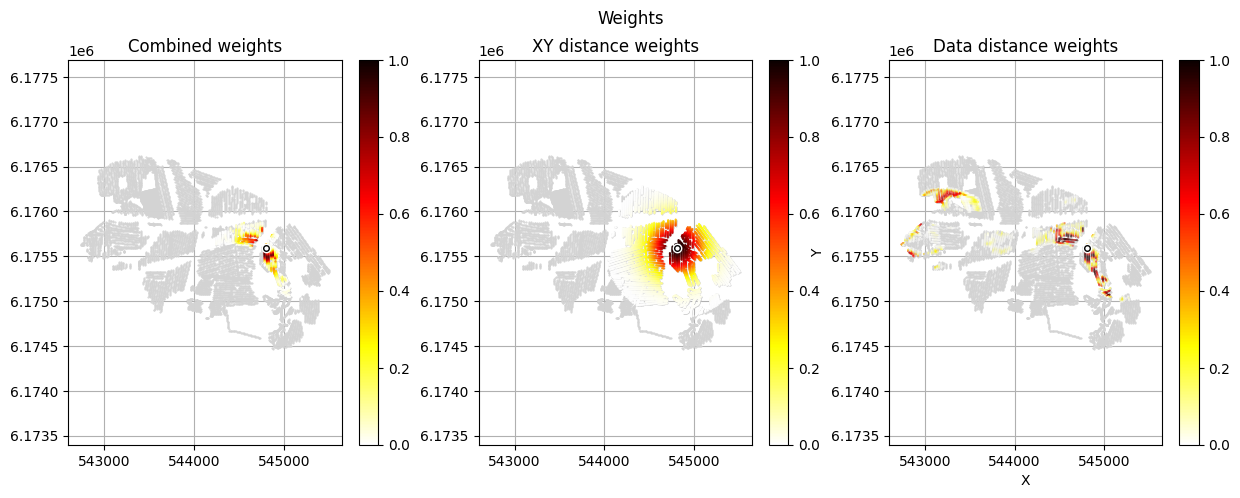
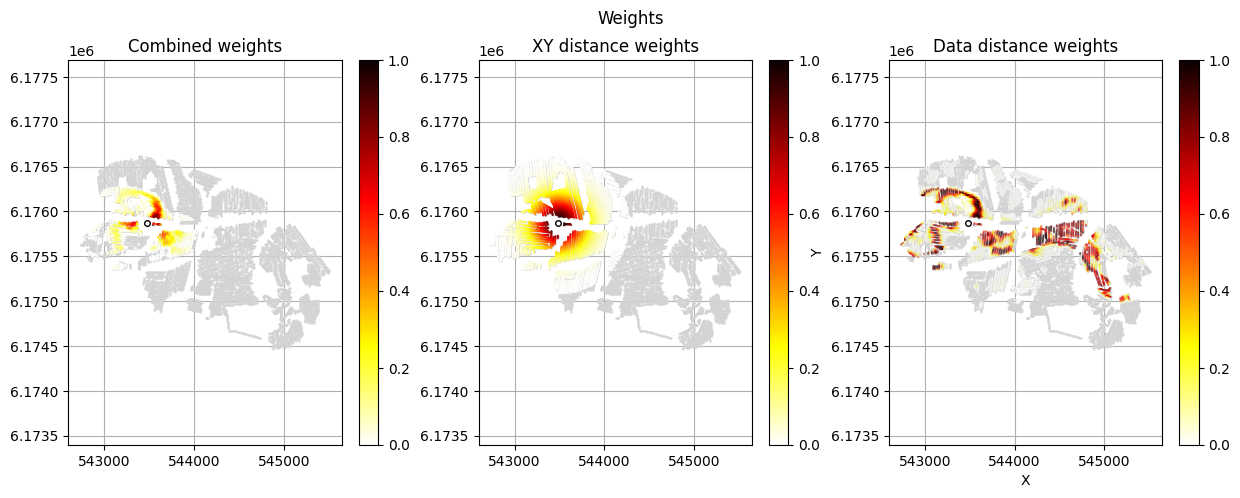
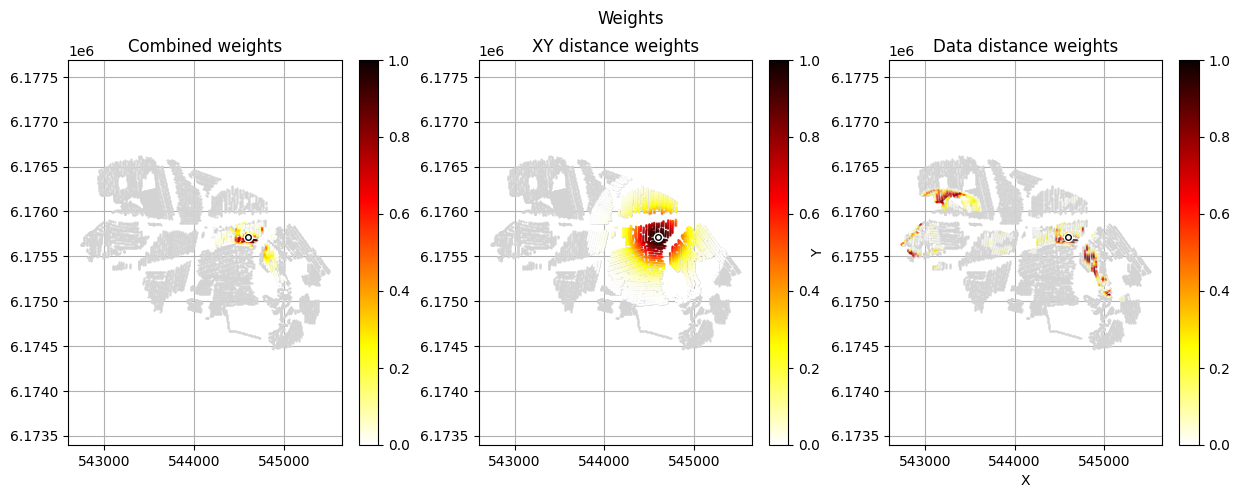
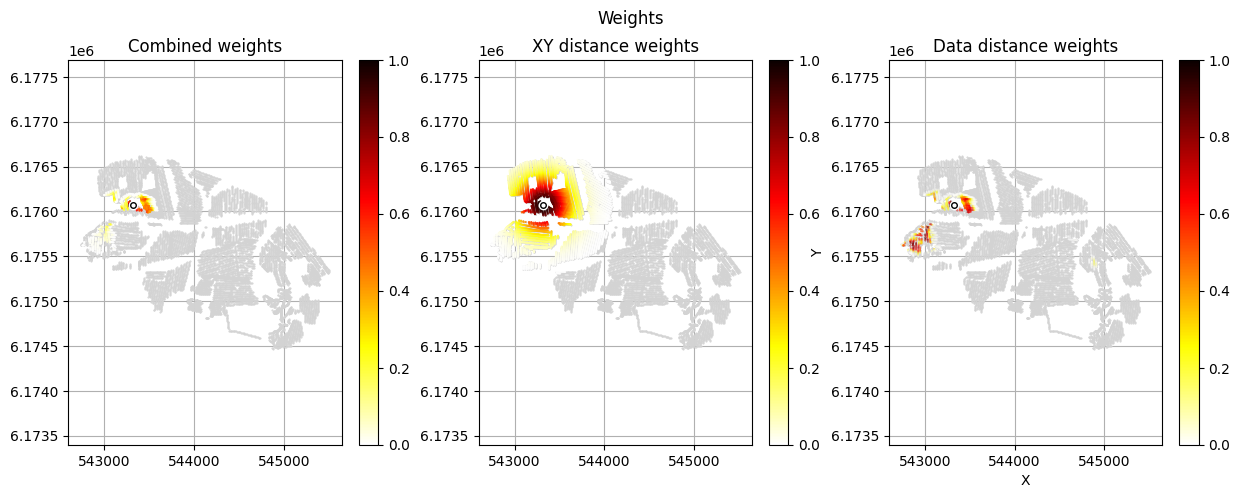
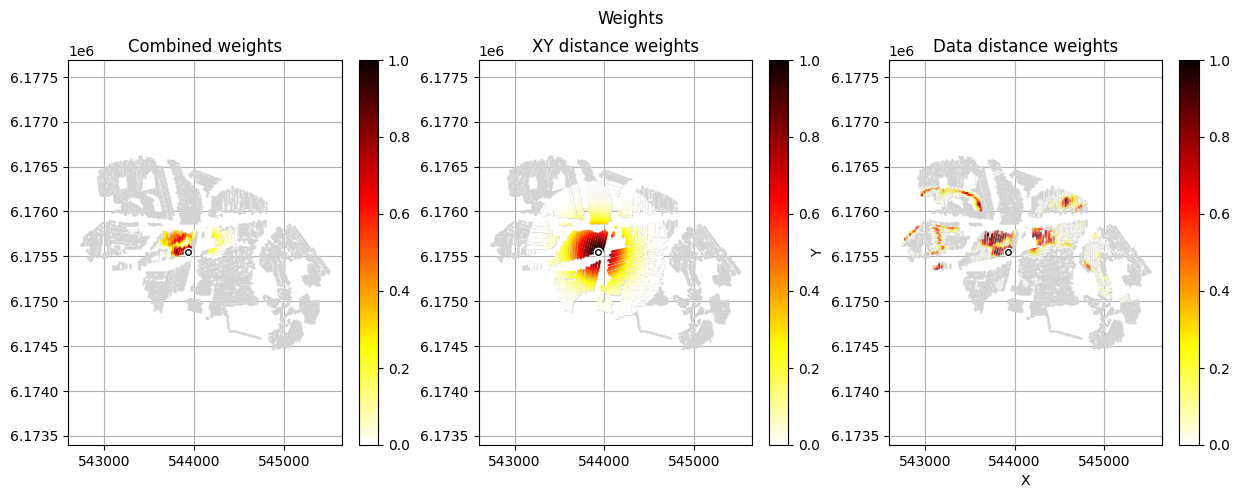
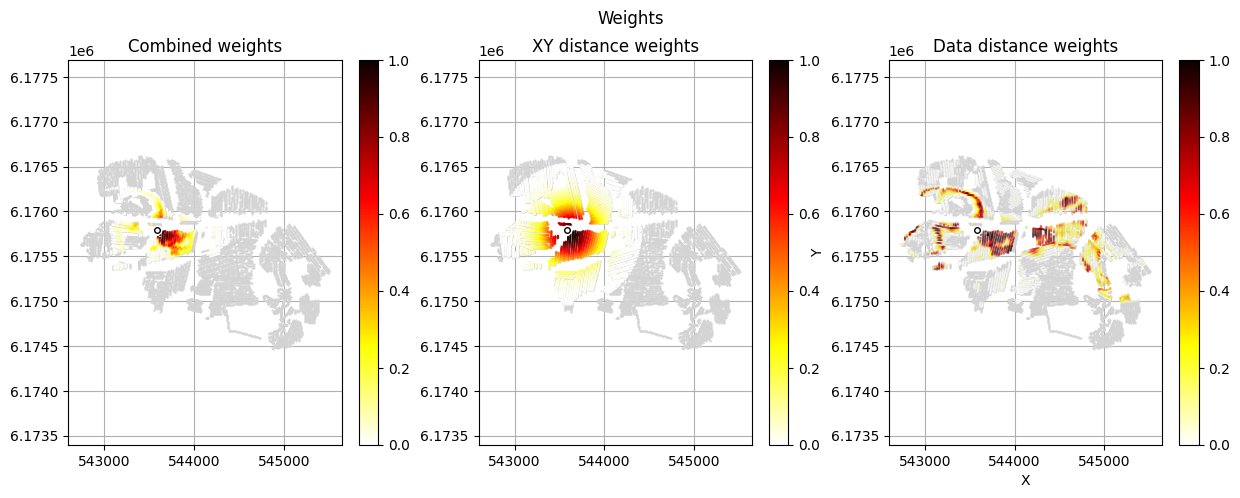
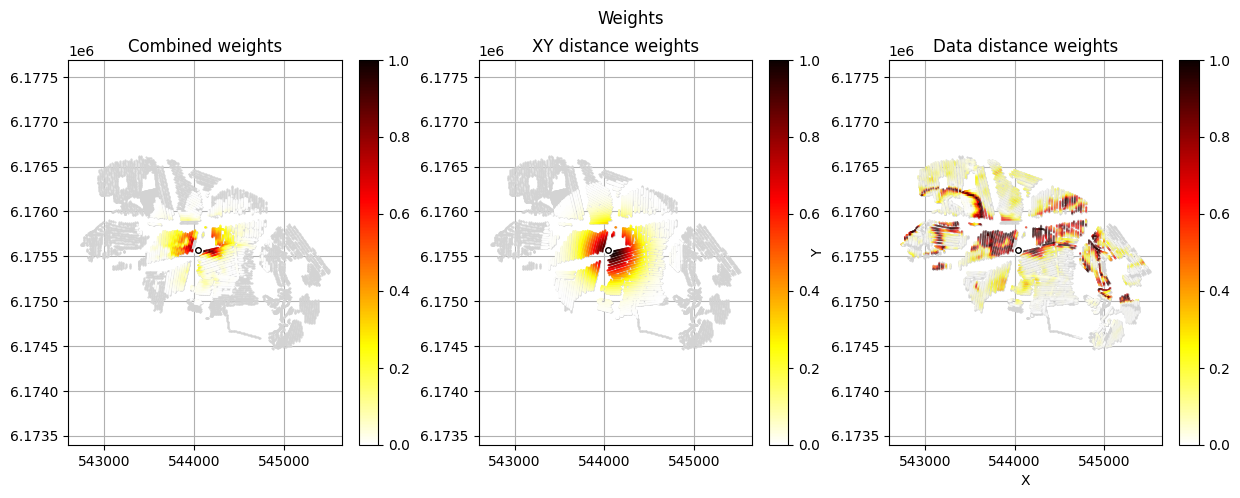
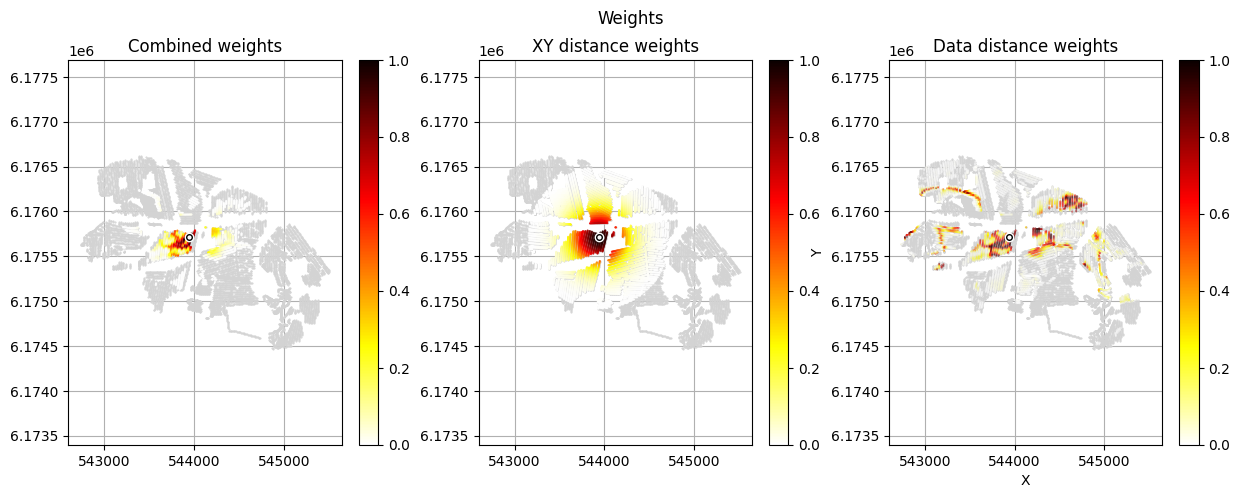
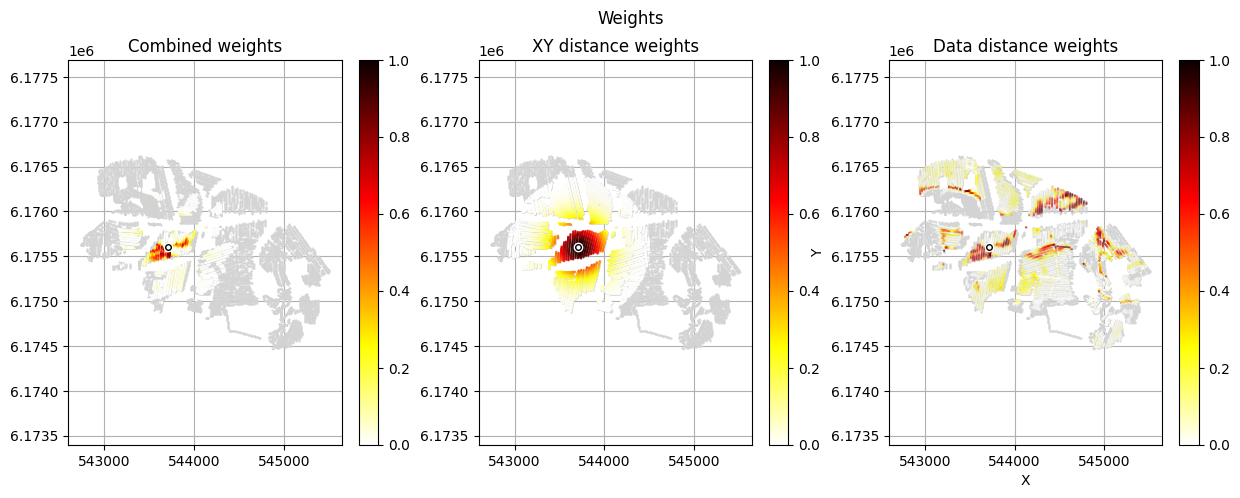
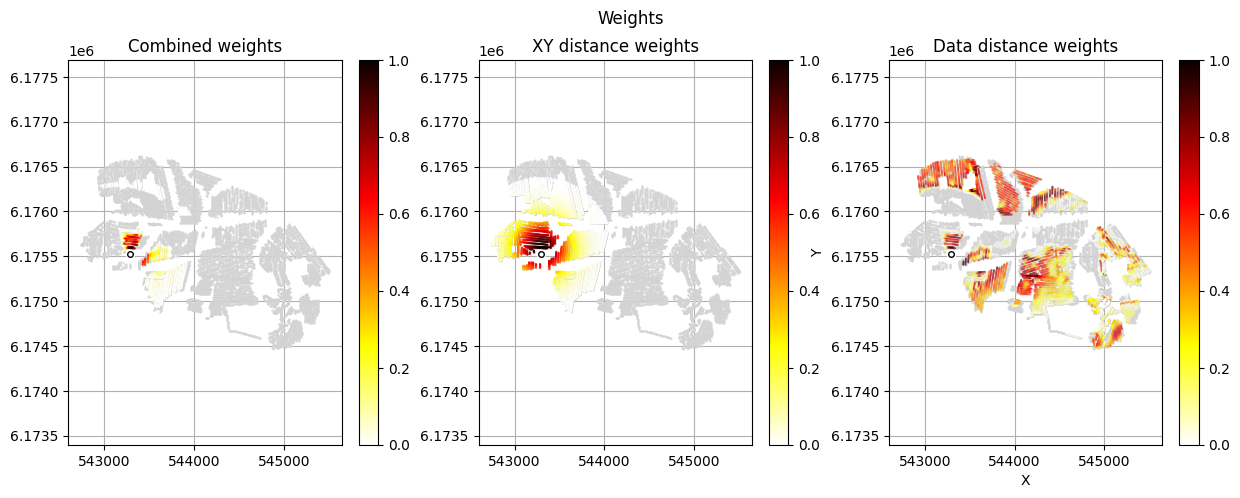
[12]:
#
# For each borehole BH in BHOLES:
# 1. Compute prior borehole data (mode class per interval per realization)
# and save to f_prior_h5 → returns P_obs and id_prior
# 2. Extrapolate point observations across the survey grid with
# distance-based weighting → d_obs, i_use
# 3. Save the observed borehole data to f_data_h5 → id_out
im_prior = 2 # lithology model index (M2)
r_data = 4 # tTEMN data based radius (db/dT) — If this is very high it will have no effect
r_dis = 300 # fade-out radius (m) — weight approaches zero at this distance
# Compute and save prior + observed data for all boreholes in one step per borehole
id_borehole_list = []
for BH in BHOLES:
id_prior, id_out = ig.save_borehole_data(
f_prior_h5, f_data_h5, BH,
im_prior=im_prior, r_data=r_data, r_dis=r_dis,
doPlot=True,
showInfo=1)
id_borehole_list.append(id_out)
prior_data_borehole: borehole=DAU01 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 5 intervals
/mnt/space/space_au11687/PROGRAMMING/integrate_module/integrate/integrate_borehole.py:757: RuntimeWarning: invalid value encountered in log10
d_test = np.log10(d_obs[:,i_use])
Using id=2
Trying to write D2 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D2
save_borehole_data: DAU01 → prior D2, data D2
prior_data_borehole: borehole=DAU02 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 6 intervals
Using id=3
Trying to write D3 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D3
save_borehole_data: DAU02 → prior D3, data D3
prior_data_borehole: borehole=DAU03 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 8 intervals
Using id=4
Trying to write D4 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D4
save_borehole_data: DAU03 → prior D4, data D4
prior_data_borehole: borehole=DAU04 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 10 intervals
Using id=5
Trying to write D5 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D5
save_borehole_data: DAU04 → prior D5, data D5
prior_data_borehole: borehole=DAU05 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 4 intervals
Using id=6
Trying to write D6 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D6
save_borehole_data: DAU05 → prior D6, data D6
prior_data_borehole: borehole=DAU06 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 3 intervals
Using id=7
Trying to write D7 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D7
save_borehole_data: DAU06 → prior D7, data D7
prior_data_borehole: borehole=DAU07 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 5 intervals
Using id=8
Trying to write D8 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D8
save_borehole_data: DAU07 → prior D8, data D8
prior_data_borehole: borehole=116. 1602 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 7 intervals
Using id=9
Trying to write D9 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D9
save_borehole_data: 116. 1602 → prior D9, data D9
prior_data_borehole: borehole=116. 535 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 3 intervals
Using id=10
Trying to write D10 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D10
save_borehole_data: 116. 535 → prior D10, data D10
prior_data_borehole: borehole=116. 1866 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 12 intervals
Using id=11
Trying to write D11 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D11
save_borehole_data: 116. 1866 → prior D11, data D11
prior_data_borehole: borehole=116. 1867 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 4 intervals
Using id=12
Trying to write D12 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D12
save_borehole_data: 116. 1867 → prior D12, data D12
prior_data_borehole: borehole=116. 1868 method=mode_probability
compute_P_obs_sparse: Processing 200000 realizations, 2 intervals
Using id=13
Trying to write D13 to tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
Adding group tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5:D13
save_borehole_data: 116. 1868 → prior D13, data D13
Select a profile¶
[13]:
X, Y, LINE, ELEVATION = ig.get_geometry(f_data_h5)
# find the index of the [X,Y] points closts to the two boreholes
i_bh = []
for i, BH in enumerate(BHOLES):
d = np.sqrt((X-BH['X'])**2 + (Y-BH['Y'])**2)
i_closest = np.argmin(d)
print("Closest point to %s is at index %d with distance %.1f m" % (BH['name'], i_closest, d[i_closest]))
i_bh.append(i_closest)
# Find points within buffer distance
X1 = 542983.01
Y1 = 6175822.76
X2 = 543584.098
Y2 = 6175788.478
Xl = np.array([X1-100,X1, X2, X2+1500])
Yl = np.array([Y1, Y1, Y2, Y2-150])
buffer = 15.0
indices, distances, segment_ids = ig.find_points_along_line_segments(
X, Y, Xl, Yl, tolerance=buffer
)
id_line = indices
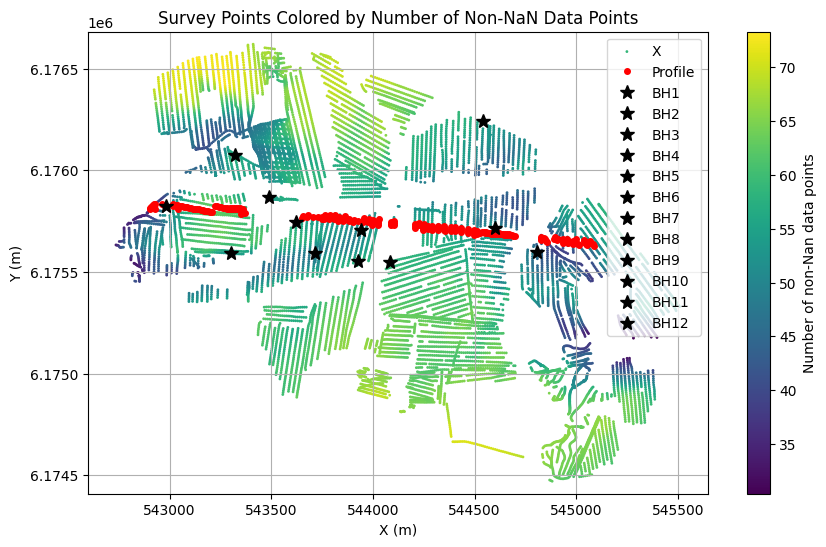
plt.figure(figsize=(10, 6))
plt.scatter(X, Y, c=ELEVATION, s=1,label='X')
plt.plot(X[id_line],Y[id_line], 'r.', markersize=8, label='Profile', zorder=2, linewidth=5)
for i in range(len(i_bh)):
plt.plot(X[i_bh[i]], Y[i_bh[i]], 'k*', markersize=10, label='BH%d' % (i+1), zorder=3)
plt.grid()
plt.colorbar(label='Number of non-Nan data points')
plt.xlabel('X (m)')
plt.ylabel('Y (m)')
plt.title('Survey Points Colored by Number of Non-NaN Data Points')
plt.axis('equal')
plt.legend()
if hardcopy:
plt.savefig('DAUGAARD_survey_points_nonnan.png', dpi=300)
plt.show()
i1=np.min(id_line)
i2=np.max(id_line)+1
Closest point to DAU01 is at index 9147 with distance 53.6 m
Closest point to DAU02 is at index 6343 with distance 0.0 m
Closest point to DAU03 is at index 5658 with distance 0.5 m
Closest point to DAU04 is at index 11552 with distance 12.5 m
Closest point to DAU05 is at index 9222 with distance 0.2 m
Closest point to DAU06 is at index 6027 with distance 0.3 m
Closest point to DAU07 is at index 7851 with distance 0.5 m
Closest point to 116. 1602 is at index 7406 with distance 58.0 m
Closest point to 116. 535 is at index 3305 with distance 45.9 m
Closest point to 116. 1866 is at index 7585 with distance 5.0 m
Closest point to 116. 1867 is at index 7520 with distance 5.0 m
Closest point to 116. 1868 is at index 7213 with distance 62.9 m
INVERSION¶
The data is now ready for inversion with the rejection sampler.
On total we have 3 data types (one tTEM and two WellLog). They can be all jointly inverted (the default) or one can select which data types to ínver using id_use
id_use = [1] # tTEM
id_use = [2] # Well 1
id_use = [3] # Well 2
id_use = [2,3] # Wells 1,2
id_use = [1,2,3] # tTEM, Wells 1,2 (the default if id_use is not set)
id_use = [1,2,3,4,5,6,7,8,9,10,11,12,13] # tTEM, and all 12 borehole data channels (the default if id_use is not set)
[14]:
# This prt of the can be rerun using different selection of data types without rerunning the abobe parts
nr=1000
T_N_above=50
T_P_acc_level=0.2
autoT = 1 # We need minium of T_N_above realizations with an acceptance probability above T_P_acc_level
id_use_arr = []
id_use_arr.append([1]) # tTEM
#id_use_arr.append([2]) # Well 1
#id_use_arr.append([3]) # Well 2
#id_use_arr.append([2,3]) # Well 1,2
#id_use_arr.append([2,3]) # Well 1,2
#id_use_arr.append([1,2,3]) # tTEM, Well 1,2
id_use_arr.append([2,3,4,5,6,7,8,9,10,11,12,13]) # All 12 borehole data channels
id_use_arr.append([1,2,3,4,5,6,7,8,9,10,11,12,13]) # All 12 borehole data channels + tTEM
N_use = N
f_post_h5_list = []
for id_use in id_use_arr:
# get string from id_use
fileparts = os.path.splitext(f_data_h5)
id_use_str = '_'.join(map(str, id_use))
f_post_h5 = 'post_%s_id%s.h5' % (fileparts[0], id_use_str)
f_post_h5 = ig.integrate_rejection(f_prior_h5,
f_data_h5,
f_post_h5,
showInfo=1,
N_use = N_use,
id_use = id_use,
nr=nr,
T_N_above = T_N_above,
T_P_acc_level = T_P_acc_level,
updatePostStat=True)
f_post_h5_list.append(f_post_h5)
Loading data from tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5. Using data types: [1]
- D1: id_prior=1, gaussian, Using 11687/40 data
Loading prior data from daugaard_merged_prior_N100000.h5. Using prior data ids: [1]
- /D1: N,nd = 100000/40
<--INTEGRATE_REJECTION-->
f_prior_h5=daugaard_merged_prior_N100000.h5, f_data_h5=tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
f_post_h5=post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
integrate_rejection: Time= 87.8s/11687 soundings, 7.5ms/sounding, 133.1it/s. T_av=35.9, EV_av=-39.5
Computing posterior statistics for 11687 of 11687 data points
Computing number of unique realizations (N_UNIQUE)
Updating /N_UNIQUE in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M1/Mean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M1/Median in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M1/Std in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M1/LogMean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M2/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M2/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M2/P
Creating /M3/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M3/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1.h5
Creating /M3/P
Loading data from tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5. Using data types: [2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13]
- D2: id_prior=2, multinomial, Using 11687/8 data
- D3: id_prior=3, multinomial, Using 11687/8 data
- D4: id_prior=4, multinomial, Using 11687/8 data
- D5: id_prior=5, multinomial, Using 11687/8 data
- D6: id_prior=6, multinomial, Using 11687/8 data
- D7: id_prior=7, multinomial, Using 11687/8 data
- D8: id_prior=8, multinomial, Using 11687/8 data
- D9: id_prior=9, multinomial, Using 11687/8 data
- D10: id_prior=10, multinomial, Using 11687/8 data
- D11: id_prior=11, multinomial, Using 11687/8 data
- D12: id_prior=12, multinomial, Using 11687/8 data
- D13: id_prior=13, multinomial, Using 11687/8 data
Loading prior data from daugaard_merged_prior_N100000.h5. Using prior data ids: [np.int64(2), np.int64(3), np.int64(4), np.int64(5), np.int64(6), np.int64(7), np.int64(8), np.int64(9), np.int64(10), np.int64(11), np.int64(12), np.int64(13)]
- /D2: N,nd = 100000/5
- /D3: N,nd = 100000/6
- /D4: N,nd = 100000/8
- /D5: N,nd = 100000/10
- /D6: N,nd = 100000/4
- /D7: N,nd = 100000/3
- /D8: N,nd = 100000/5
- /D9: N,nd = 100000/7
- /D10: N,nd = 100000/3
- /D11: N,nd = 100000/12
- /D12: N,nd = 100000/4
- /D13: N,nd = 100000/2
<--INTEGRATE_REJECTION-->
f_prior_h5=daugaard_merged_prior_N100000.h5, f_data_h5=tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
f_post_h5=post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
integrate_rejection: Time= 19.0s/11687 soundings, 1.6ms/sounding, 614.4it/s. T_av=1.1, EV_av=-11.5
Computing posterior statistics for 11687 of 11687 data points
Computing number of unique realizations (N_UNIQUE)
Updating /N_UNIQUE in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Mean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Median in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Std in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/LogMean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/P
Creating /M3/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M3/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M3/P
Loading data from tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5. Using data types: [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13]
- D1: id_prior=1, gaussian, Using 11687/40 data
- D2: id_prior=2, multinomial, Using 11687/8 data
- D3: id_prior=3, multinomial, Using 11687/8 data
- D4: id_prior=4, multinomial, Using 11687/8 data
- D5: id_prior=5, multinomial, Using 11687/8 data
- D6: id_prior=6, multinomial, Using 11687/8 data
- D7: id_prior=7, multinomial, Using 11687/8 data
- D8: id_prior=8, multinomial, Using 11687/8 data
- D9: id_prior=9, multinomial, Using 11687/8 data
- D10: id_prior=10, multinomial, Using 11687/8 data
- D11: id_prior=11, multinomial, Using 11687/8 data
- D12: id_prior=12, multinomial, Using 11687/8 data
- D13: id_prior=13, multinomial, Using 11687/8 data
Loading prior data from daugaard_merged_prior_N100000.h5. Using prior data ids: [1, np.int64(2), np.int64(3), np.int64(4), np.int64(5), np.int64(6), np.int64(7), np.int64(8), np.int64(9), np.int64(10), np.int64(11), np.int64(12), np.int64(13)]
- /D1: N,nd = 100000/40
- /D2: N,nd = 100000/5
- /D3: N,nd = 100000/6
- /D4: N,nd = 100000/8
- /D5: N,nd = 100000/10
- /D6: N,nd = 100000/4
- /D7: N,nd = 100000/3
- /D8: N,nd = 100000/5
- /D9: N,nd = 100000/7
- /D10: N,nd = 100000/3
- /D11: N,nd = 100000/12
- /D12: N,nd = 100000/4
- /D13: N,nd = 100000/2
<--INTEGRATE_REJECTION-->
f_prior_h5=daugaard_merged_prior_N100000.h5, f_data_h5=tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2.h5
f_post_h5=post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
integrate_rejection: Time=100.1s/11687 soundings, 8.6ms/sounding, 116.8it/s. T_av=35.8, EV_av=-50.6
Computing posterior statistics for 11687 of 11687 data points
Computing number of unique realizations (N_UNIQUE)
Updating /N_UNIQUE in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Mean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Median in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/Std in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M1/LogMean in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M2/P
Creating /M3/Mode in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M3/Entropy in post_tTEM_20230727_AVG_export_tTEM_20230814_AVG_export_tTEM_20230829_AVG_export_tTEM_20230913_AVG_export_tTEM_20231109_AVG_export_WF_gf2_id1_2_3_4_5_6_7_8_9_10_11_12_13.h5
Creating /M3/P
POSTERIOR ANALYSIS¶
[15]:
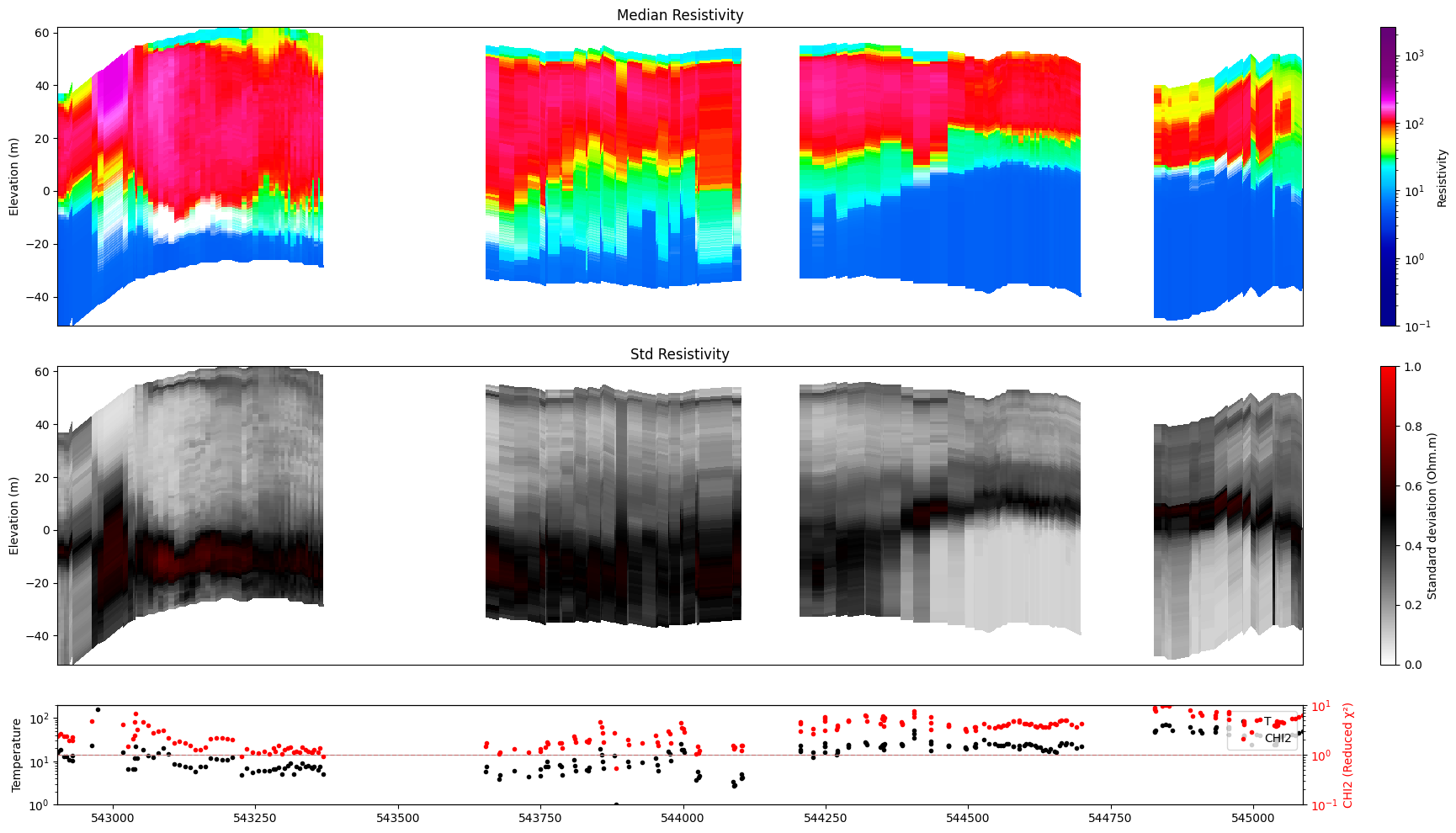
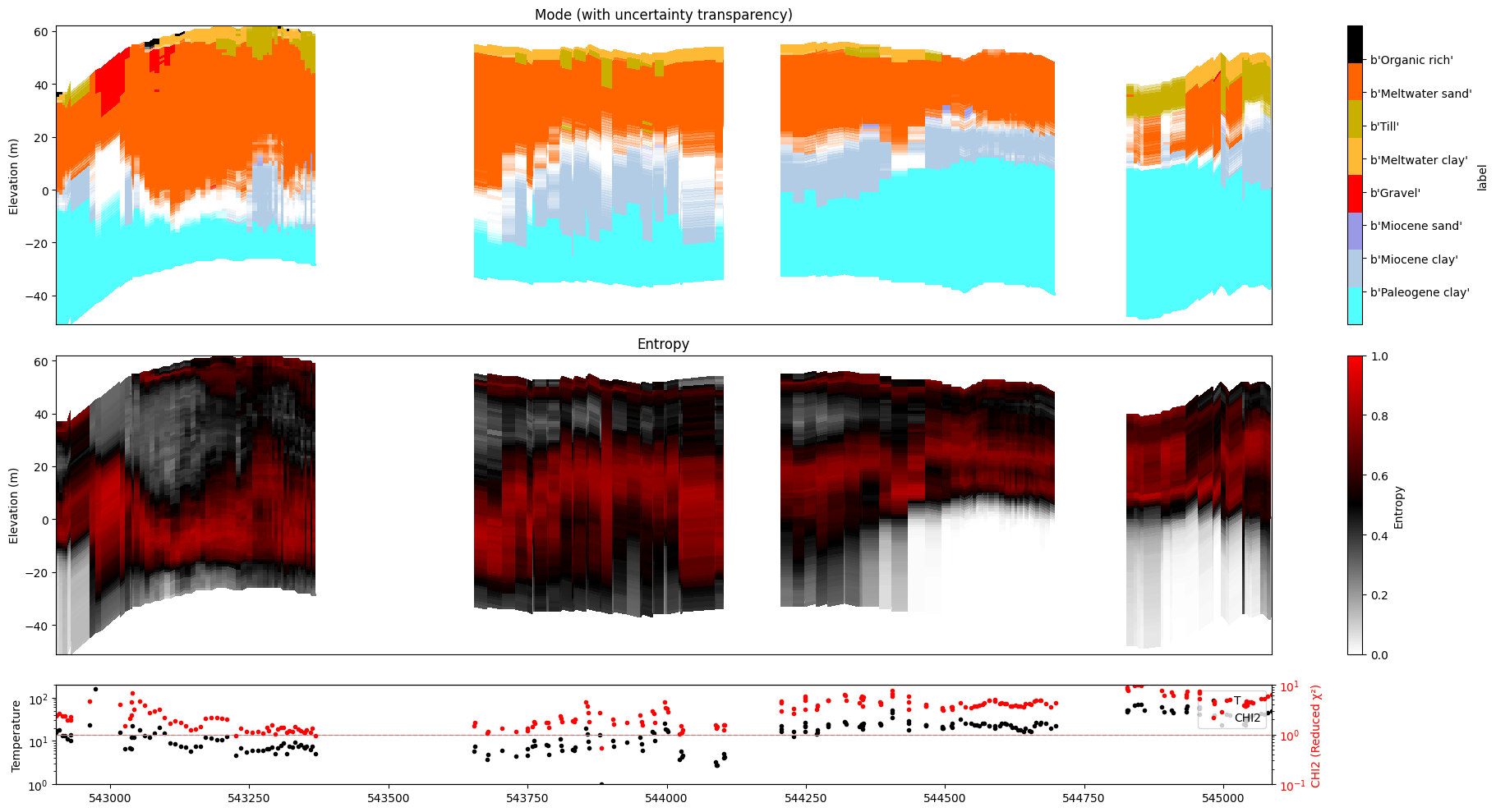
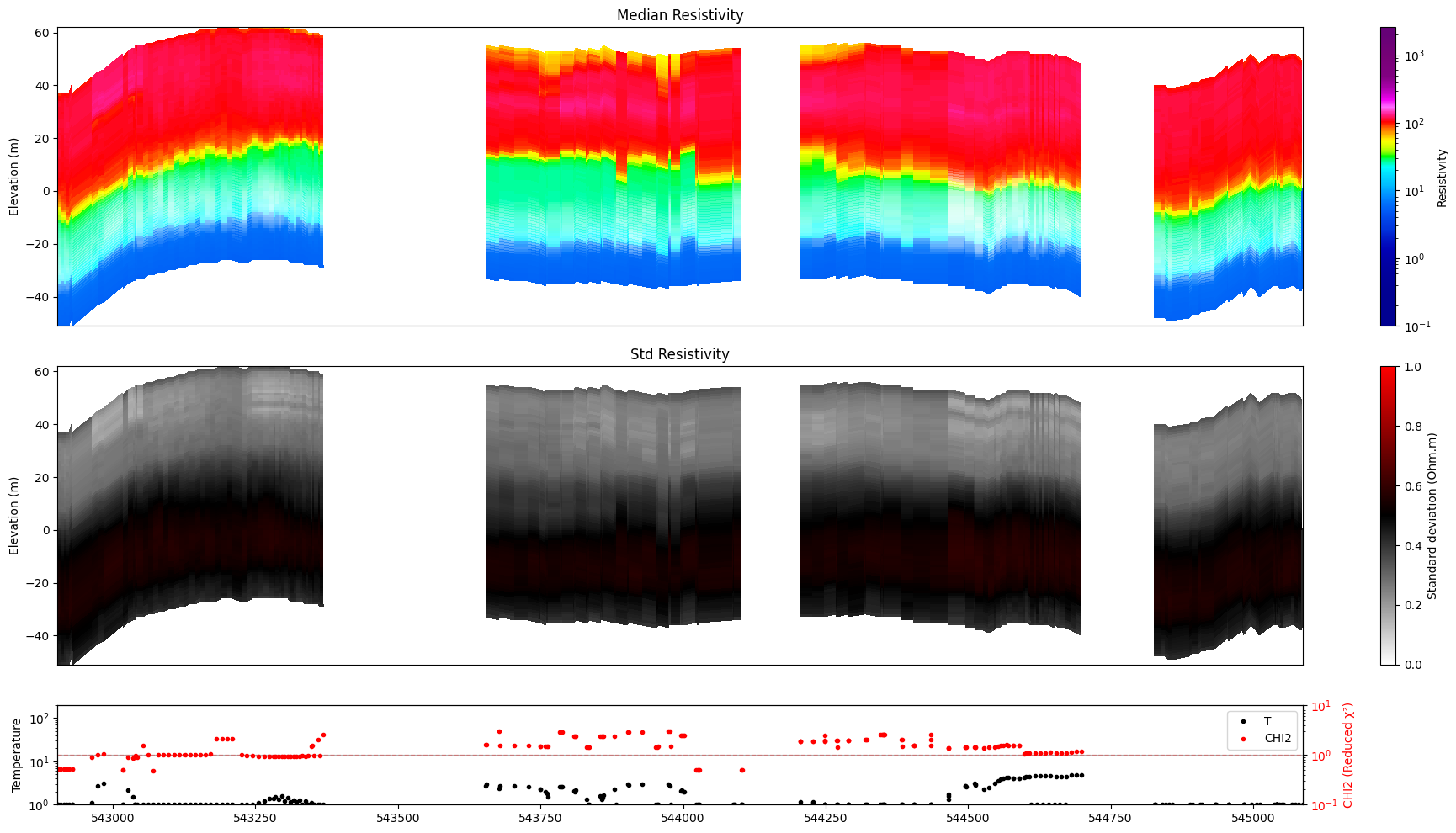
for f_post_h5 in f_post_h5_list:
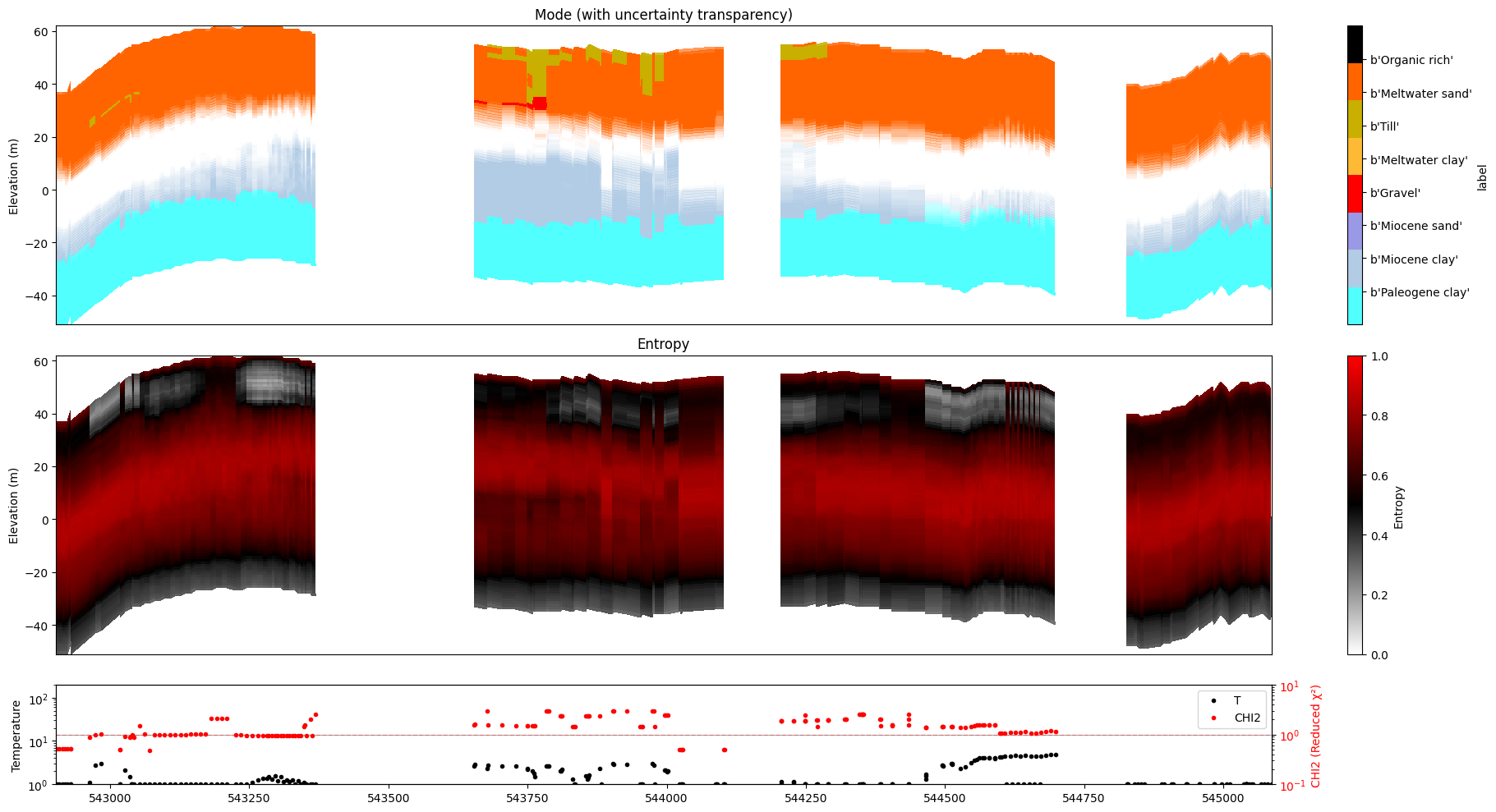
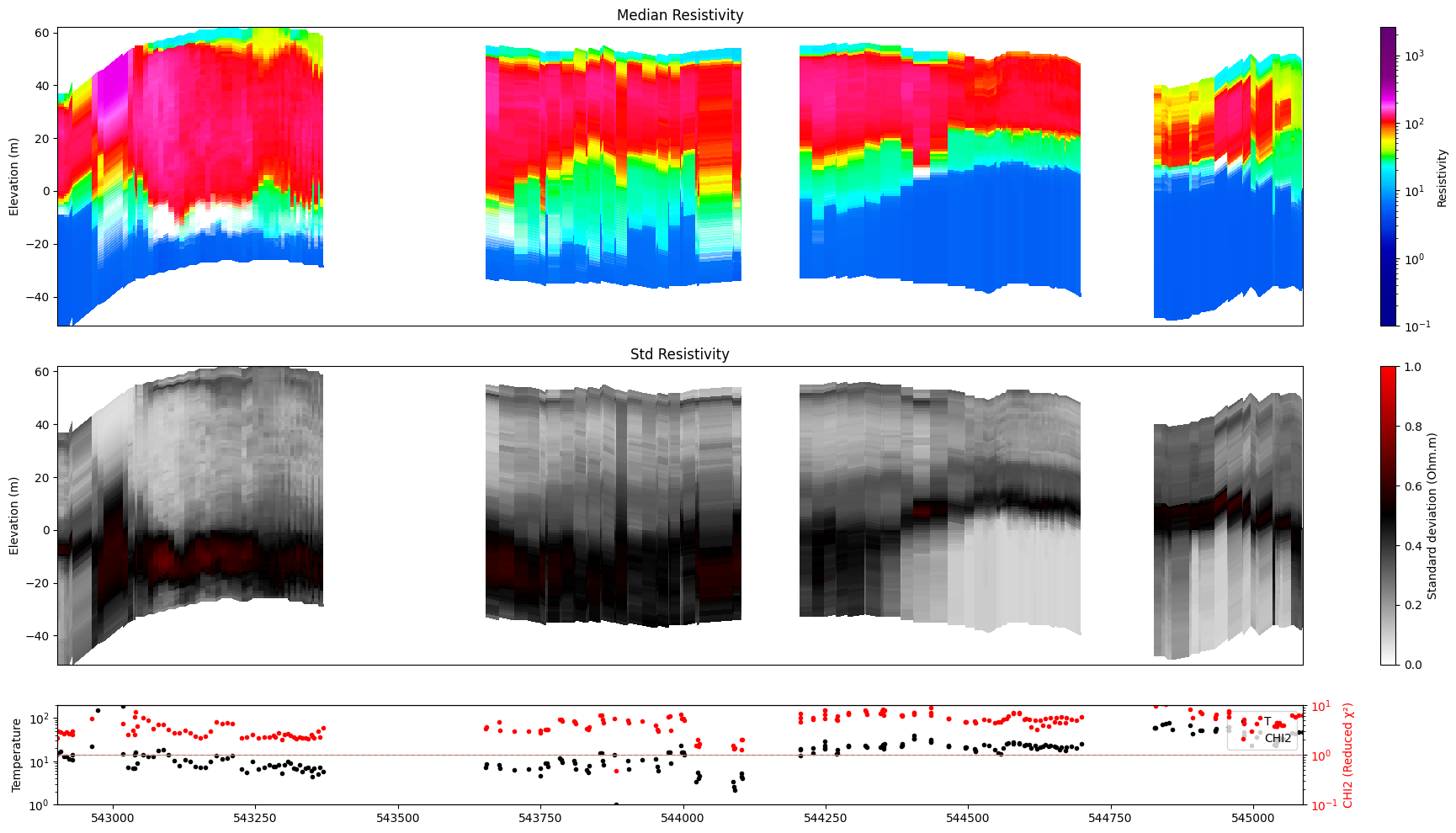
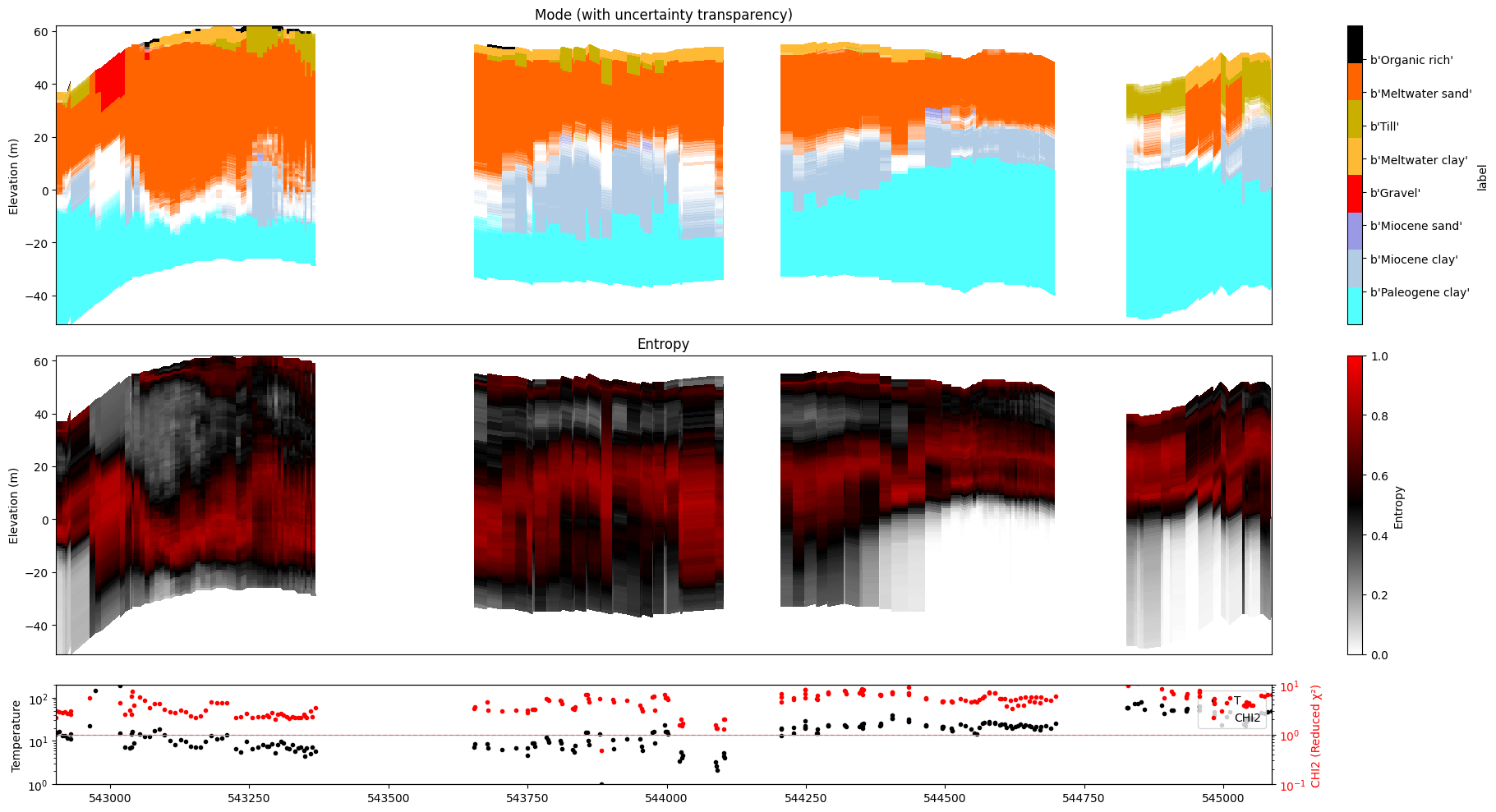
ig.plot_profile(f_post_h5, im=1, ii=id_line, gap_threshold=100, xaxis='x', hardcopy=hardcopy, alpha = 1,std_min = 0.5, std_max = 0.6)
ig.plot_profile(f_post_h5, im=2, ii=id_line, gap_threshold=100, xaxis='x', hardcopy=hardcopy, alpha=1, entropy_min =0.7, entropy_max=0.8)
Getting cmap from attribute
0.0
1.0
Getting cmap from attribute
0.0105724335
1.0
Getting cmap from attribute
0.0
1.0
[16]:
for f_post_h5 in f_post_h5_list:
ig.plot_feature_2d(f_post_h5, key='Median', im=1, elevation=40)
plt.plot(X[i_bh], Y[i_bh], 'k*', markersize=10, label='Boreholes')
plt.legend()
plt.show()
/M1/Median[elev=40.0m] Resistivity
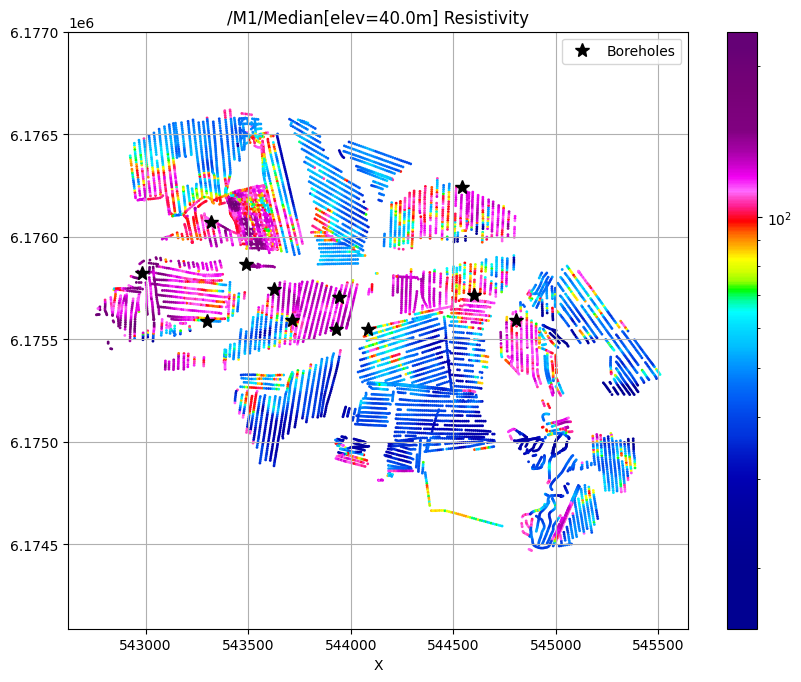
/M1/Median[elev=40.0m] Resistivity
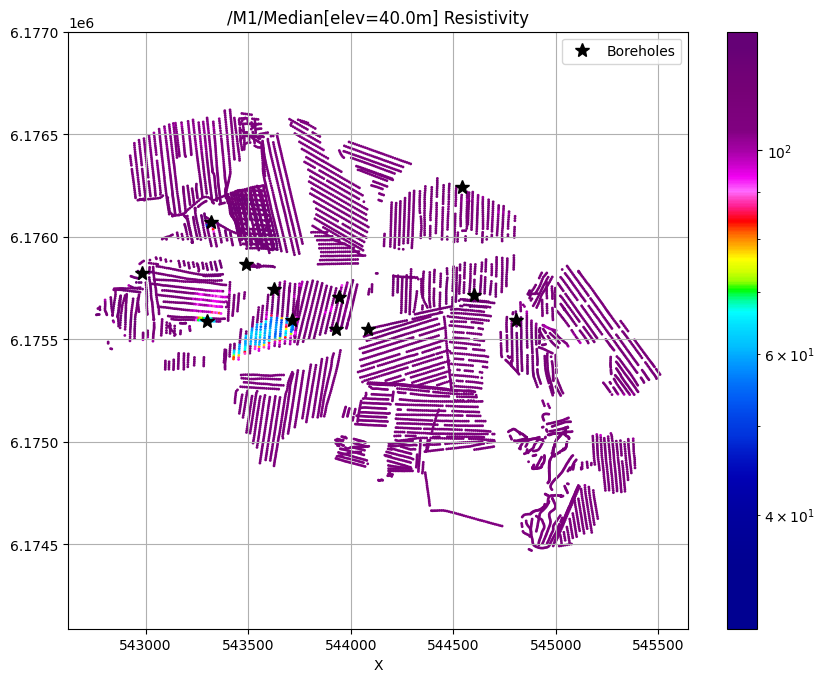
/M1/Median[elev=40.0m] Resistivity
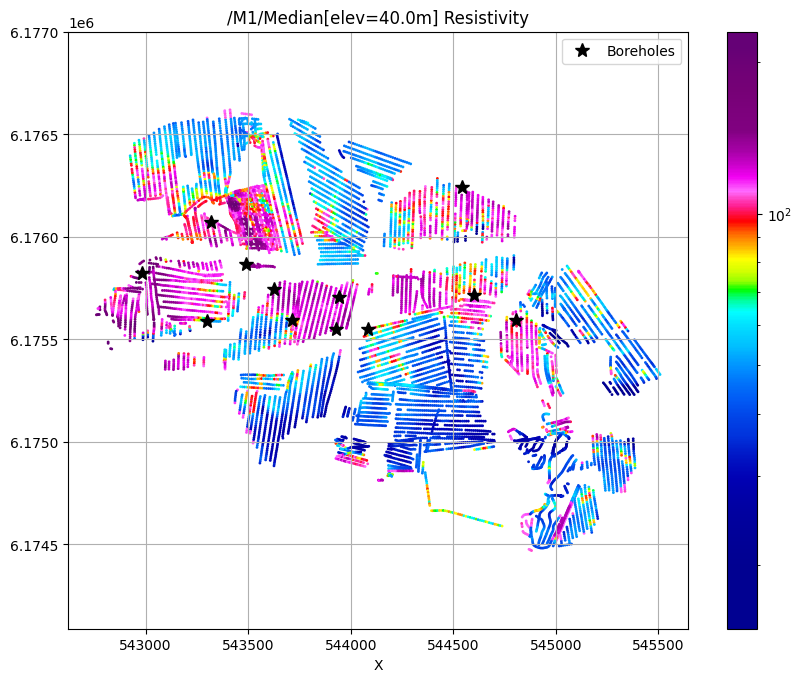
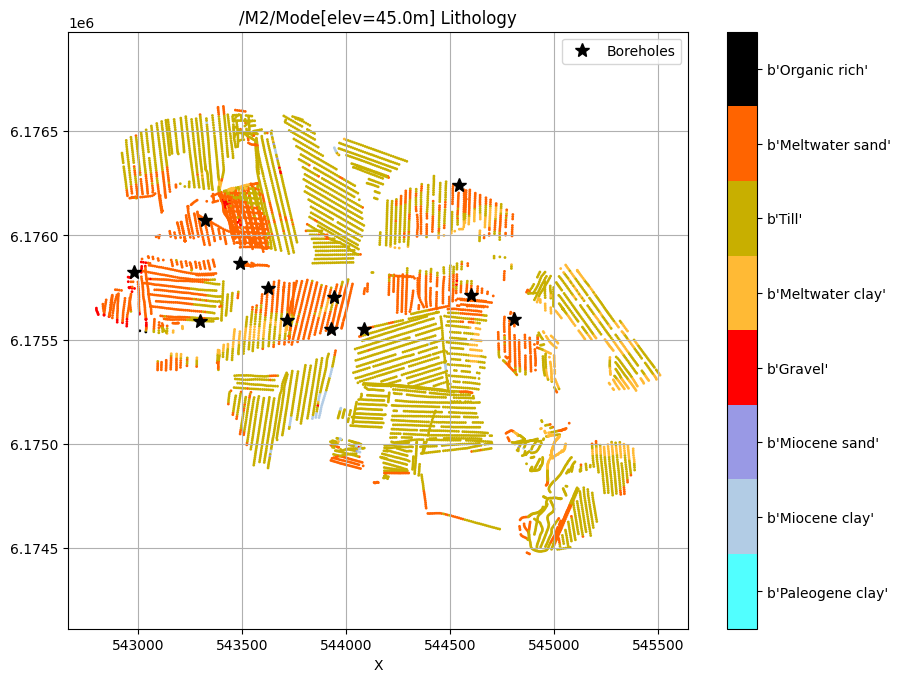
[17]:
for f_post_h5 in f_post_h5_list:
ig.plot_feature_2d(f_post_h5, key='Mode', im=2, elevation=45)
plt.plot(X[i_bh], Y[i_bh], 'k*', markersize=10, label='Boreholes')
plt.legend()
plt.show()
/M2/Mode[elev=45.0m] Lithology
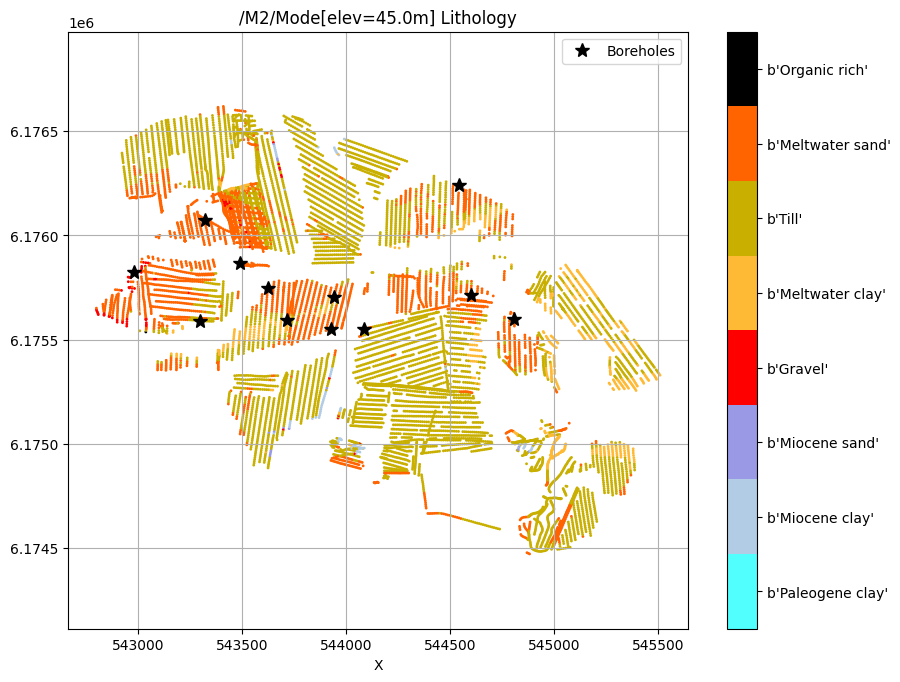
/M2/Mode[elev=45.0m] Lithology
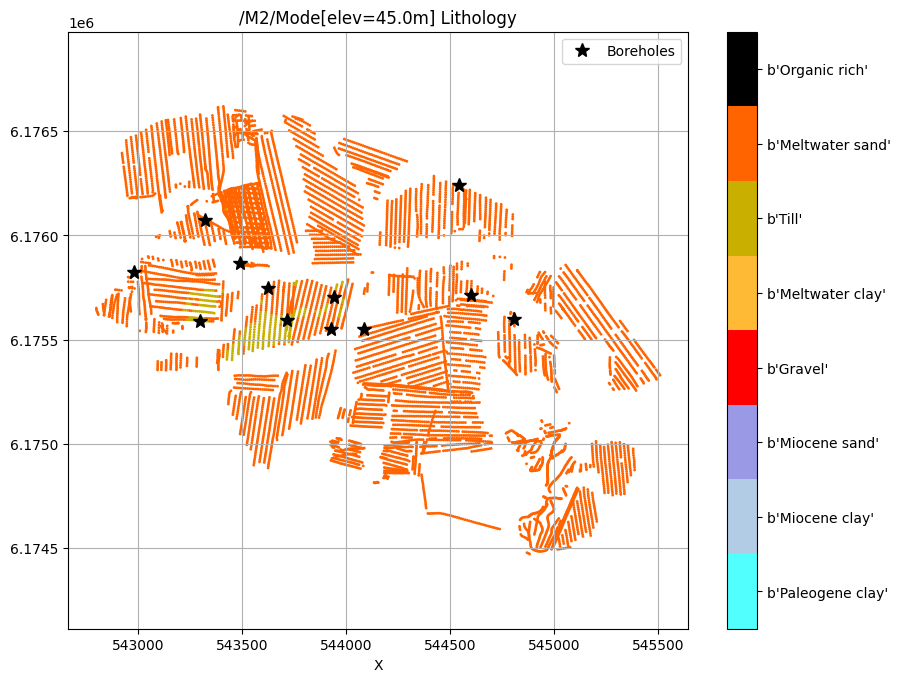
/M2/Mode[elev=45.0m] Lithology
Posterior Probability of INSIDE vs OUTSIDE¶
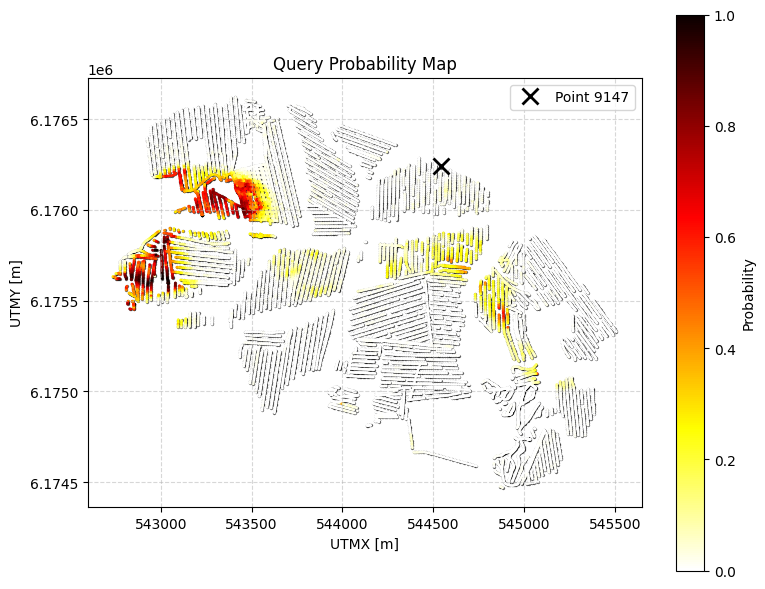
QUERY : Probability that the cumulative thickness of lithology class 2 within 0–30 m depth is greater than 10 m, and with an additional constraint: any top layer that is NOT sand/gravel cannot be thicker than 3m.
[18]:
for f_post_h5 in f_post_h5_list:
ig.plot_feature_2d(f_post_h5, key='Mode', im=3, iz=0, cmap='jet')
plt.plot(X[i_bh], Y[i_bh], 'k*', markersize=10, label='Boreholes')
plt.legend()
plt.show()
/M3/Mode[0,:] Source File Index
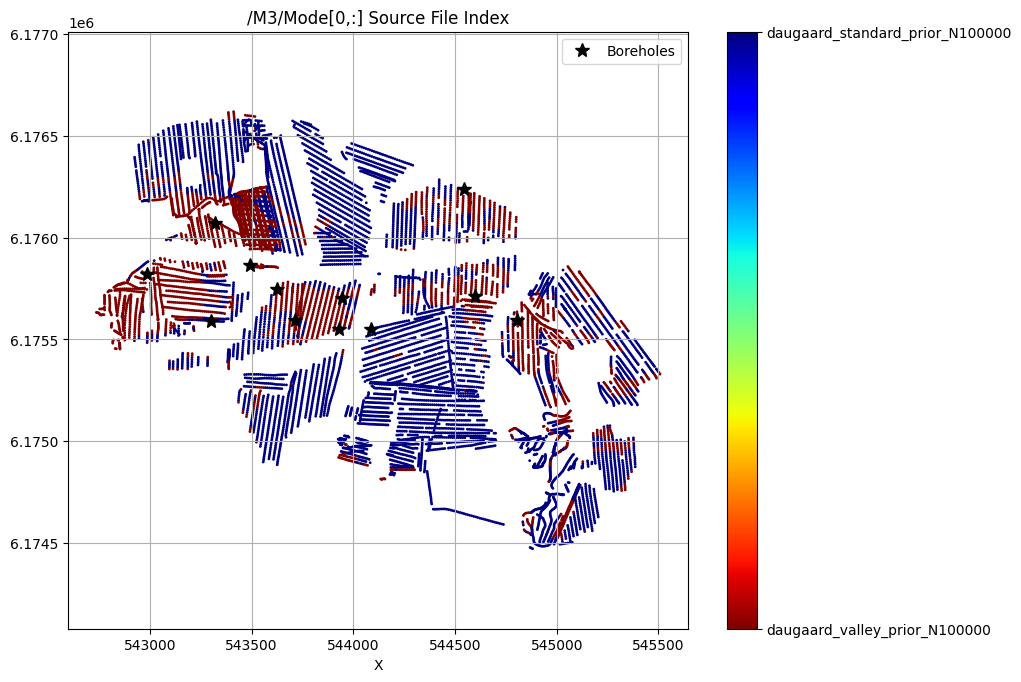
/M3/Mode[0,:] Source File Index
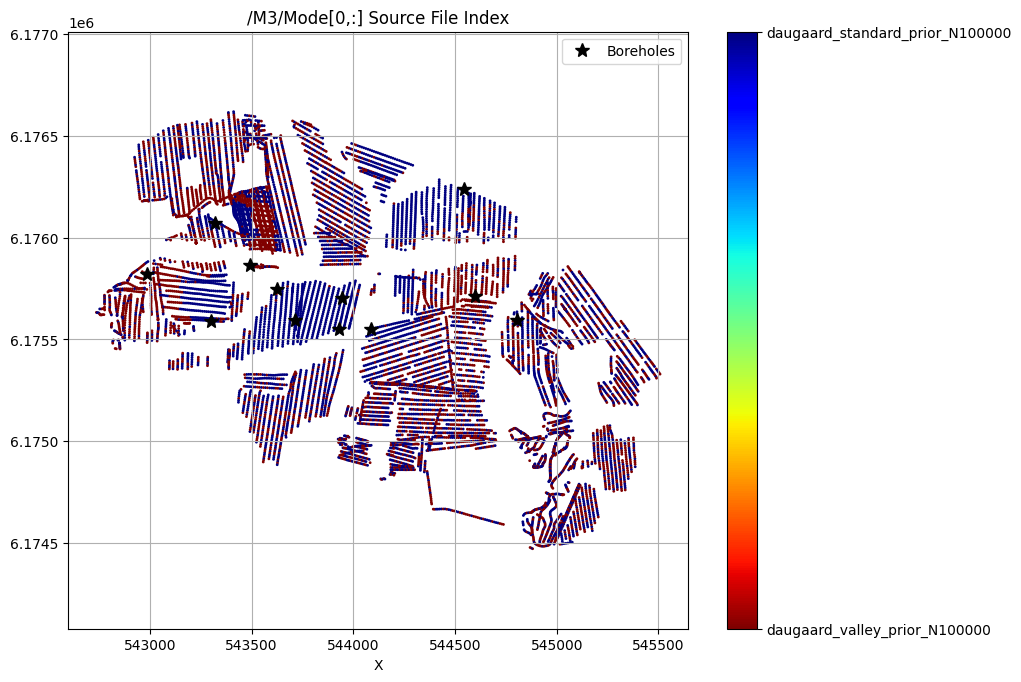
/M3/Mode[0,:] Source File Index
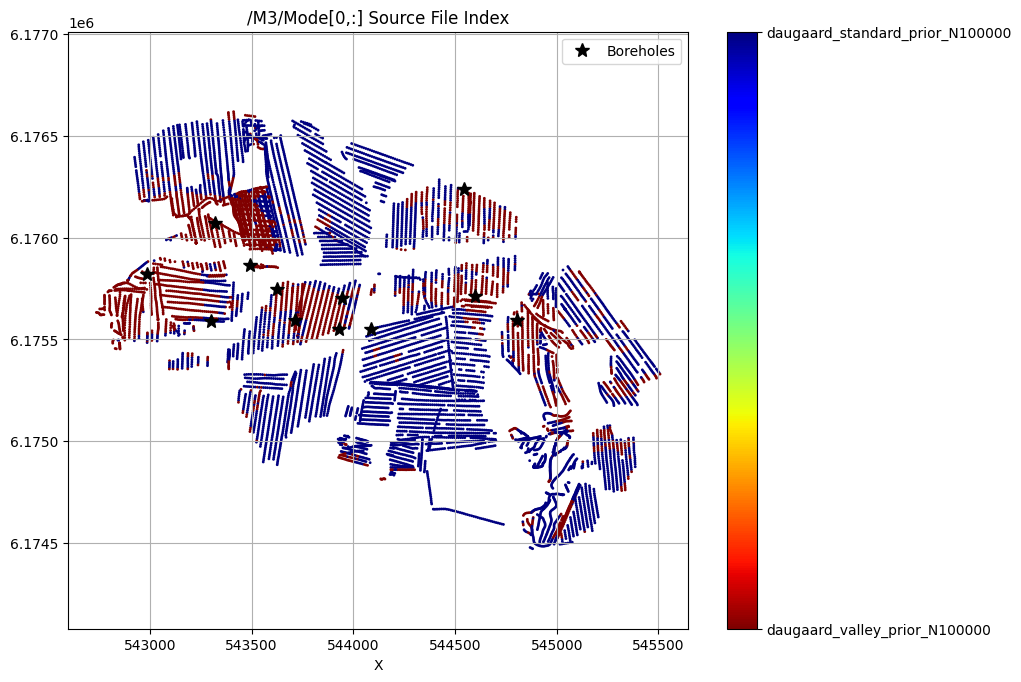
QUERY POSTERIOR MODEL REALIZATIONS¶
QUERY : Probability that the cumulative thickness of lithology class 2 within 0–30 m depth is greater than 10 m, and with an additional constraint: any top layer that is NOT sand/gravel cannot be thicker than 3m.
[ ]:
doLoadQuery = False
if doLoadQuery:
query = ig.load_query('query_ex1.json')
else:
query = {
"constraints": [
{
"im": 2,
"classes": [2, 5],
"thickness_mode": "cumulative",
"thickness_comparison": ">",
"thickness_threshold": 20.0,
"depth_min": 0.0,
"depth_max": 30.0,
"negate": False
},
{
"im": 2,
"classes": [1, 3, 4, 6, 7, 8], # All classes except sand (2) and gravel (5)
"thickness_mode": "first_occurrence",
"thickness_comparison": "<",
"thickness_threshold": 3.0,
"depth_min": 0.0,
"depth_max": 30.0,
"negate": False
}
]
}
ig.save_query(query, 'query_daugaard.json')
for f_post_h5 in f_post_h5_list:
# Compute the probability of the query being satisfied for each model realization in the posterior, and get metadata about the query results (e.g. which realizations satisfy the query, etc.)
P, meta = ig.query(f_post_h5, query)
# Plot the predicted probability map, and the the outcome at the borehole locations (which should be close to 1 since the query is based on the borehole data)
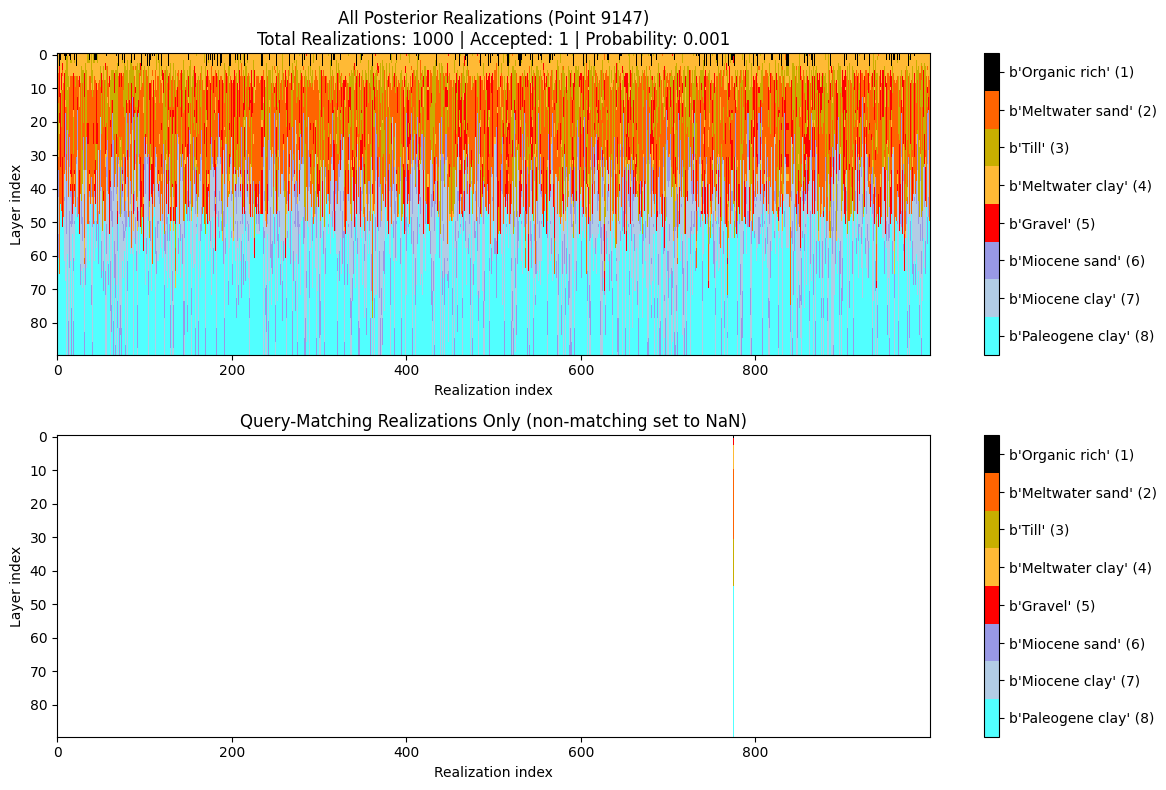
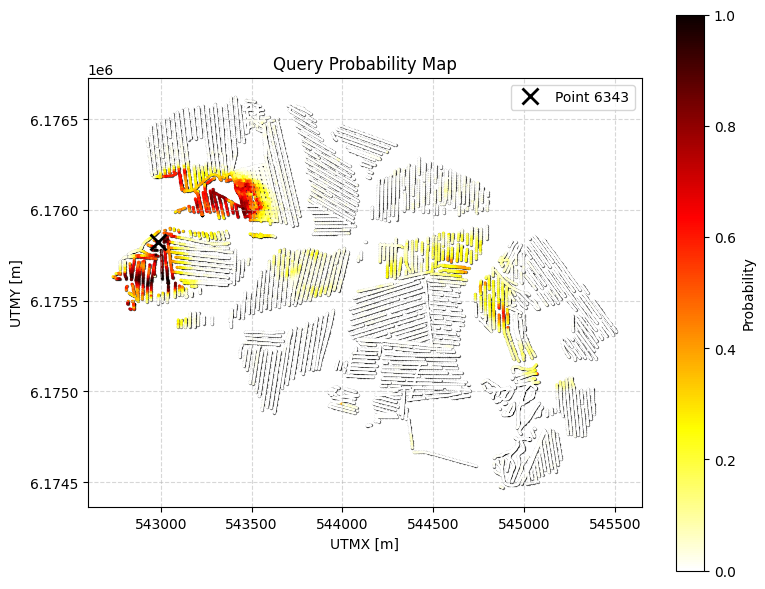
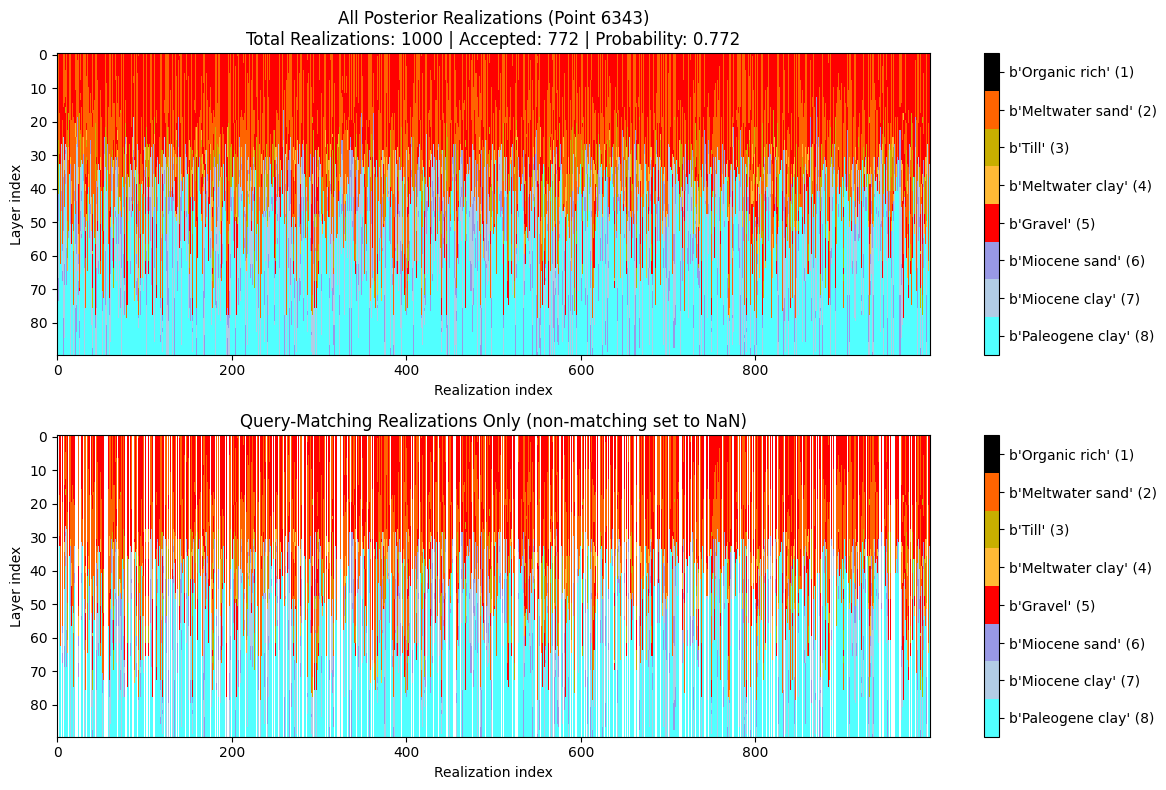
ig.query_plot(P, meta, ip=i_bh[0], query_dict=query, f_post_h5=f_post_h5)
ig.query_plot(P, meta, ip=i_bh[1], query_dict=query, f_post_h5=f_post_h5)
Query saved to query_daugaard.json
Evaluating query: 100%|████████████████████████████████████████████████████████████████████████████| 11687/11687 [00:49<00:00, 237.52location/s]
Evaluating query: 24%|██████████████████▏ | 2765/11687 [00:06<00:21, 406.04location/s]