Dual data types (low and high moment seperately)¶
[1]:
try:
# Check if the code is running in an IPython kernel (which includes Jupyter notebooks)
get_ipython()
# If the above line doesn't raise an error, it means we are in a Jupyter environment
# Execute the magic commands using IPython's run_line_magic function
get_ipython().run_line_magic('load_ext', 'autoreload')
get_ipython().run_line_magic('autoreload', '2')
except:
# If get_ipython() raises an error, we are not in a Jupyter environment
# # #%load_ext autoreload
# # #%autoreload 2
pass
[2]:
import integrate as ig
# check if parallel computations can be performed
parallel = ig.use_parallel(showInfo=1)
import h5py
import numpy as np
import matplotlib.pyplot as plt
hardcopy=True
Notebook detected. Parallel processing is OK
[3]:
case = 'DAUGAARD'
files = ig.get_case_data(case=case)
f_data_h5 = files[0]
f_data_h5 = 'DAUGAARD_AVG.h5'
file_gex= ig.get_gex_file_from_data(f_data_h5)
print("Using data file: %s" % f_data_h5)
print("Using GEX file: %s" % file_gex)
Getting data for case: DAUGAARD
--> Got data for case: DAUGAARD
Using data file: DAUGAARD_AVG.h5
Using GEX file: TX07_20231016_2x4_RC20-33.gex
1. Setup the prior model (\(\rho(\mathbf{m},\mathbf{d})\)¶
In this example a simple layered prior model will be considered
1a. first, a sample of the prior model parameters, \(\rho(\mathbf{m})\), will be generated¶
[4]:
N=25000
# Layered model
f_prior_h5 = ig.prior_model_layered(N=N,lay_dist='chi2', NLAY_deg=3, RHO_min=1, RHO_max=3000)
f_prior_data_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex, parallel=parallel, showInfo=0)
ig.integrate_update_prior_attributes(f_prior_data_h5)
File PRIOR_CHI2_NF_3_log-uniform_N25000.h5 does not exist.
Using file_basename=TX07_20231016_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 32.5s/25000 soundings. 1.3ms/sounding, 770.2it/s
[5]:
# into TWO data sets, LOW and HIGH moment.
ig.plot_data_prior(f_prior_data_h5, f_data_h5)
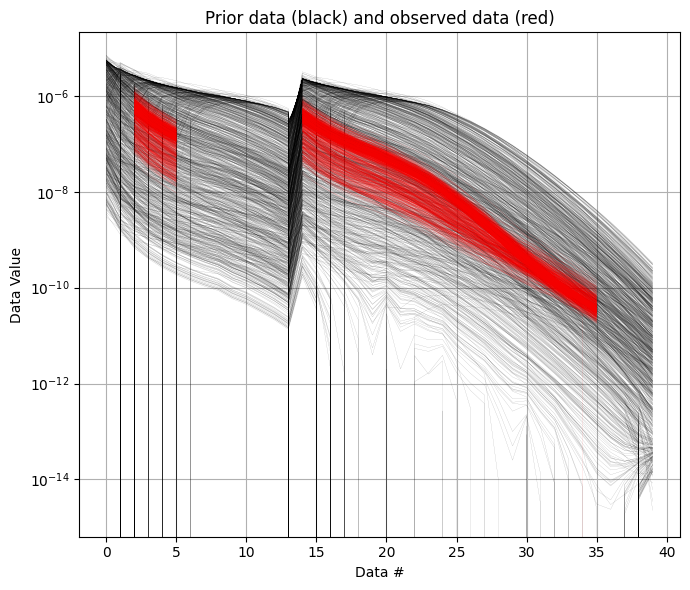
[5]:
True
[6]:
# Read D_obs from f_data_h5
with h5py.File(f_data_h5, 'r') as f:
D_obs = f['D1/d_obs'][:]
D_std = f['D1/d_std'][:]
# Alternatively, use the ig.load_data function
#D_obs = ig.load_data(f_data_h5, id=1, showInfo=1)['d_obs'][0]
#D_std = ig.load_data(f_data_h5, id=1, showInfo=1)['d_std'][0]
with h5py.File(f_prior_data_h5, 'r') as f:
D = f['/D1'][:]
# Alternatively, use the ig.load_prior_data function
#D = ig.load_prior_data(f_prior_data_h5)[0][0]
# Now splot the into low and high moment data sets
# The low moment data set will be the first 14 columns, and the high moment data set will be the last columns.
nd = D_obs.shape[1]
n_low = 14
n_high = nd - n_low
# set i low to 0:n_low-1
i_low = range(n_low)
i_high = range(n_low, nd)
# Split prior data
D_low = D[:,i_low]
D_high = D[:,i_high]
# Split observed data
D_obs_low = D_obs[:,i_low]
D_std_low = D_std[:,i_low]*2
D_obs_high = D_obs[:,i_high]
D_std_high = D_std[:,i_high]*2
plt.semilogy(D_obs[0],'k.',markersize=10, label='D_obs')
plt.semilogy(D_obs_low[0],'.', color='gray', markersize=6, label='D_obs_low')
plt.semilogy(D_obs_high[0],'.', color='gray',markersize=4, label='D_obs_high')
plt.semilogy(D[0],'b.',markersize=10, label='D_obs')
plt.semilogy(D_low[0],'.', color='yellow', markersize=6, label='D_obs_low')
plt.semilogy(D_high[0],'.', color='yellow', markersize=6, label='D_obs_low')
plt.legend()
[6]:
<matplotlib.legend.Legend at 0x774224f1d040>
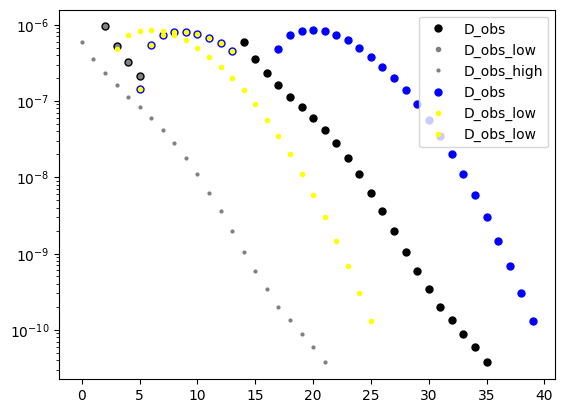
[ ]:
[7]:
f_data_dual_h5 = 'DAUGAARD_AVG_dual.h5'
useOldMethod = False
if useOldMethod:
ig.copy_hdf5_file(f_data_h5,f_data_dual_h5)
# Delete D1
with h5py.File(f_data_dual_h5, 'a') as f:
# show groups in f['']
if 'D1' in f.keys():
del(f['D1'])
# Update D1 and D2
with h5py.File(f_data_dual_h5, 'a') as f:
# remove 'D1'
#del f['D1']
f.create_dataset('D1/d_obs', data=D_obs_low)
f.create_dataset('D1/d_std', data=D_std_low)
f['D1'].attrs['noise_model'] = 'gaussian'
f.create_dataset('D2/d_obs', data=D_obs_high)
f.create_dataset('D2/d_std', data=D_std_high)
f['D2'].attrs['noise_model'] = 'gaussian'
else:
# Alternatively, use the ig.save_data_gaussian function
ig.copy_hdf5_file(f_data_h5,f_data_dual_h5)
ig.save_data_gaussian(D_obs_low, D_std = D_std_low, f_data_h5 = f_data_dual_h5, id=1, showInfo=0)
ig.save_data_gaussian(D_obs_high, D_std = D_std_high, f_data_h5 = f_data_dual_h5, id=2, showInfo=0)
Data has 11693 stations and 14 channels
Removing group DAUGAARD_AVG_dual.h5:D1
Adding group DAUGAARD_AVG_dual.h5:D1
Data has 11693 stations and 26 channels
Adding group DAUGAARD_AVG_dual.h5:D2
[8]:
f_prior_data_dual_h5 = 'PRIOR_dual.h5'
ig.copy_hdf5_file(f_prior_data_h5,f_prior_data_dual_h5)
if useOldMethod:
with h5py.File(f_prior_data_dual_h5, 'a') as f:
# show groups in f['']
if 'D1' in f.keys():
del(f['D1'])
f.create_dataset('D1', data=D_low)
f.create_dataset('D2', data=D_high)
else:
# Alternatively, use the ig.save_data_gaussian function
ig.save_prior_data(f_prior_data_dual_h5, D_low, id=1, force_delete=True)
ig.save_prior_data(f_prior_data_dual_h5, D_high, id=2, force_delete=False)
Deleting prior data '/D1' from file: <HDF5 file "PRIOR_dual.h5" (mode r+)>
[9]:
ig.plot_data_prior(f_prior_data_dual_h5, f_data_dual_h5, id=1)
ig.plot_data_prior(f_prior_data_dual_h5, f_data_dual_h5, id=2)
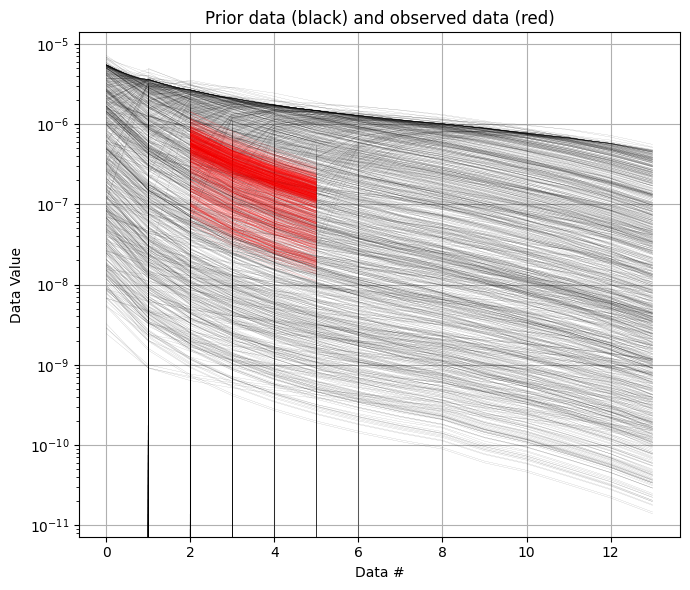
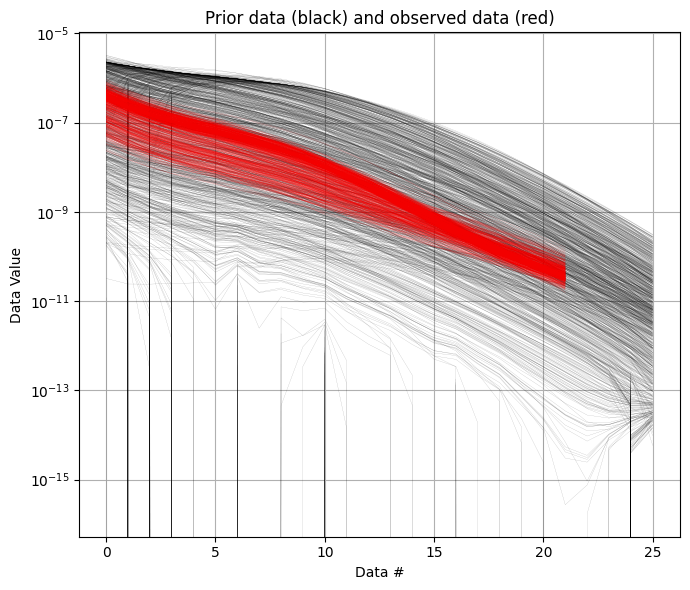
[9]:
True
Sample the posterior \(\sigma(\mathbf{m})\)¶
The posterior distribution is sampling using the extended rejection sampler.
[10]:
N_use = 100000 #%N
N_cpu = 8
f_post_arr = []
updatePostStat=False
showInfo = 1
import time
t_inversion = []
for itype in [0,1,2,3]:
t_start = time.time()
if itype == 0:
# LOW AND HIGH MOMENT AS ONE DATA SET - THE ORIGINAL METHOD
f_post_h5 = ig.integrate_rejection(f_prior_data_h5,
f_data_h5,
f_post_h5='POST_type%d.h5' % itype,
N_use = N_use, Ncpu=N_cpu,
showInfo=showInfo,
updatePostStat=updatePostStat)
elif itype == 1:
# LOW MOMENT ONLY
f_post_h5 = ig.integrate_rejection(f_prior_data_dual_h5,
f_data_dual_h5,
f_post_h5='POST_type%d.h5' % itype,
N_use = N_use, Ncpu=N_cpu,
showInfo=showInfo,
updatePostStat=updatePostStat,
id_use = [1])
elif itype == 2:
# HIGH MOMENT ONLY
f_post_h5 = ig.integrate_rejection(f_prior_data_dual_h5,
f_data_dual_h5,
# f_post_h5='POST_type%d.h5' % itype,
N_use = N_use, Ncpu=N_cpu,
showInfo=showInfo,
updatePostStat=updatePostStat,
id_use = [2])
elif itype == 3:
# JOINT INVERSION USING BOTH LOW AND HIGH MOMENT
f_post_h5 = ig.integrate_rejection(f_prior_data_dual_h5,
f_data_dual_h5,
f_post_h5='POST_type%d.h5' % itype,
N_use = N_use, Ncpu=N_cpu,
showInfo=showInfo,
updatePostStat=updatePostStat,
id_use = [1,2])
t_inversion.append(time.time() - t_start)
f_post_arr.append(f_post_h5)
Loading data from DAUGAARD_AVG.h5. Using data types: [1]
- D1: id_prior=1, gaussian, Using 11693/40 data
Loading prior data from PRIOR_CHI2_NF_3_log-uniform_N25000_TX07_20231016_2x4_RC20-33_Nh280_Nf12.h5. Using prior data ids: [1]
- /D1: N,nd = 25000/40
<--INTEGRATE_REJECTION-->
f_prior_h5=PRIOR_CHI2_NF_3_log-uniform_N25000_TX07_20231016_2x4_RC20-33_Nh280_Nf12.h5, f_data_h5=DAUGAARD_AVG.h5
f_post_h5=POST_type0.h5
integrate_rejection: Time= 47.0s/11693 soundings, 4.0ms/sounding, 248.8it/s. T_av=87.1, EV_av=-98.1
Loading data from DAUGAARD_AVG_dual.h5. Using data types: [1]
- D1: id_prior=1, gaussian, Using 11693/14 data
Loading prior data from PRIOR_dual.h5. Using prior data ids: [1]
- /D1: N,nd = 25000/14
<--INTEGRATE_REJECTION-->
f_prior_h5=PRIOR_dual.h5, f_data_h5=DAUGAARD_AVG_dual.h5
f_post_h5=POST_type1.h5
integrate_rejection: Time= 12.6s/11693 soundings, 1.1ms/sounding, 929.7it/s. T_av=1.0, EV_av=-6.2
Loading data from DAUGAARD_AVG_dual.h5. Using data types: [2]
- D2: id_prior=2, gaussian, Using 11693/26 data
Loading prior data from PRIOR_dual.h5. Using prior data ids: [2]
- /D2: N,nd = 25000/26
<--INTEGRATE_REJECTION-->
f_prior_h5=PRIOR_dual.h5, f_data_h5=DAUGAARD_AVG_dual.h5
f_post_h5=/mnt/space/space_au11687/PROGRAMMING/integrate_module/examples/POST_PRIOR_dual_Nu100000_aT1.h5
integrate_rejection: Time= 27.3s/11693 soundings, 2.3ms/sounding, 429.0it/s. T_av=17.4, EV_av=-24.6
Loading data from DAUGAARD_AVG_dual.h5. Using data types: [1, 2]
- D1: id_prior=1, gaussian, Using 11693/14 data
- D2: id_prior=2, gaussian, Using 11693/26 data
Loading prior data from PRIOR_dual.h5. Using prior data ids: [1, 2]
- /D1: N,nd = 25000/14
- /D2: N,nd = 25000/26
<--INTEGRATE_REJECTION-->
f_prior_h5=PRIOR_dual.h5, f_data_h5=DAUGAARD_AVG_dual.h5
f_post_h5=POST_type3.h5
integrate_rejection: Time= 37.6s/11693 soundings, 3.2ms/sounding, 311.0it/s. T_av=21.8, EV_av=-31.7
[11]:
for f_post_h5 in f_post_arr:
ig.integrate_posterior_stats(f_post_h5)
[12]:
for f_post_h5 in f_post_arr:
# % Plot Profiles
ig.plot_profile(f_post_h5, i1=1000, i2=2000, im=1, hardcopy=hardcopy)
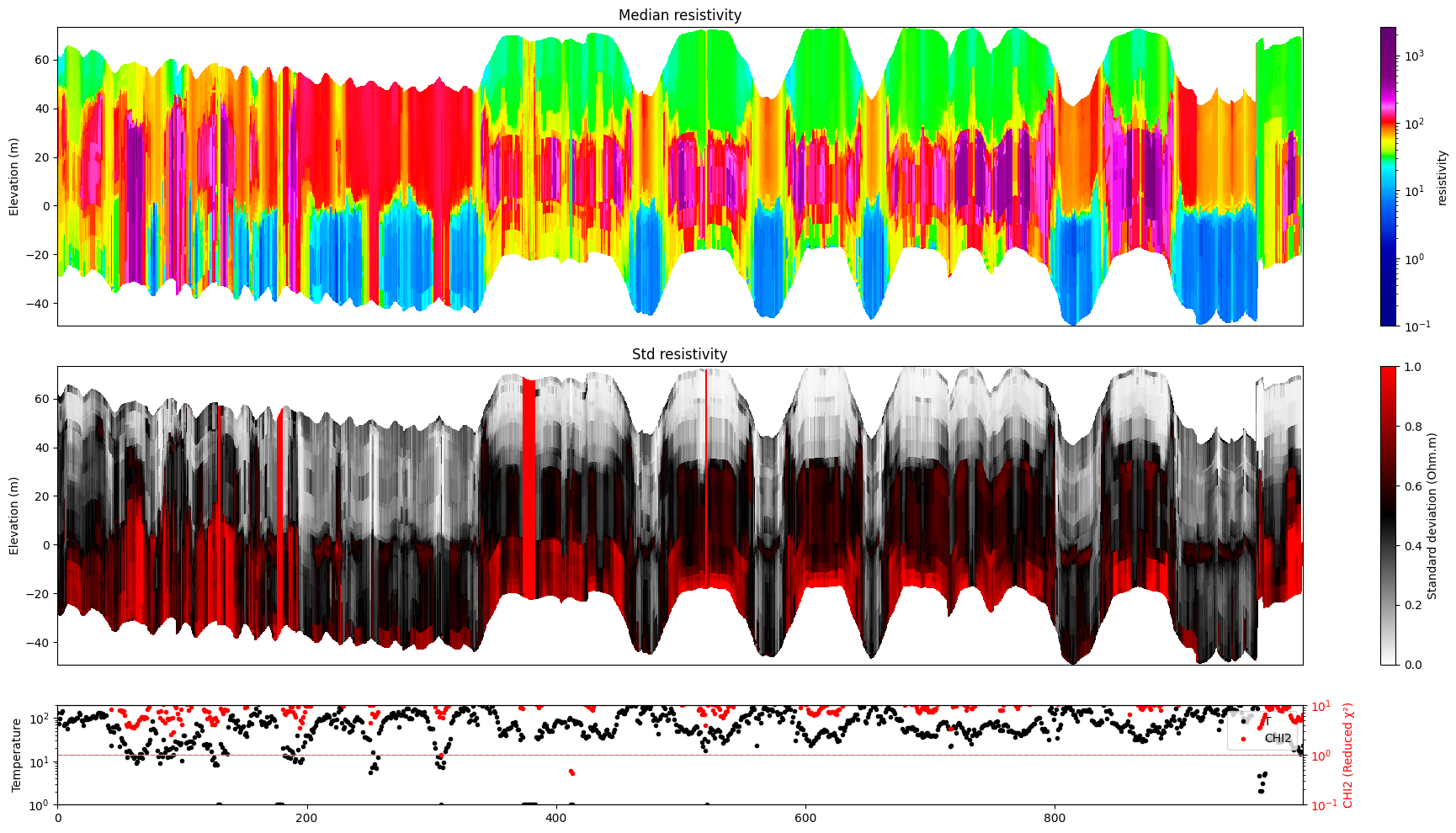
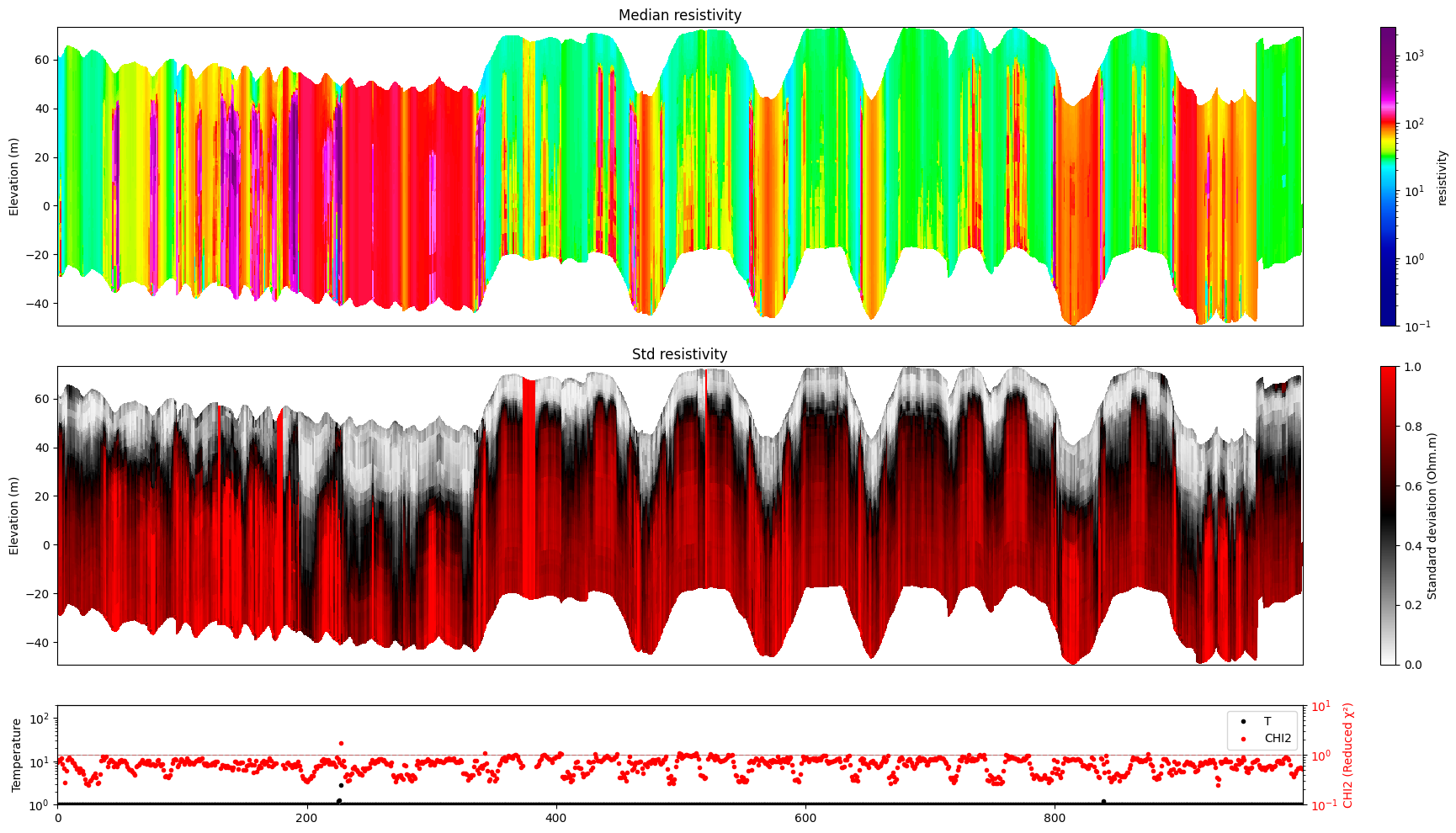
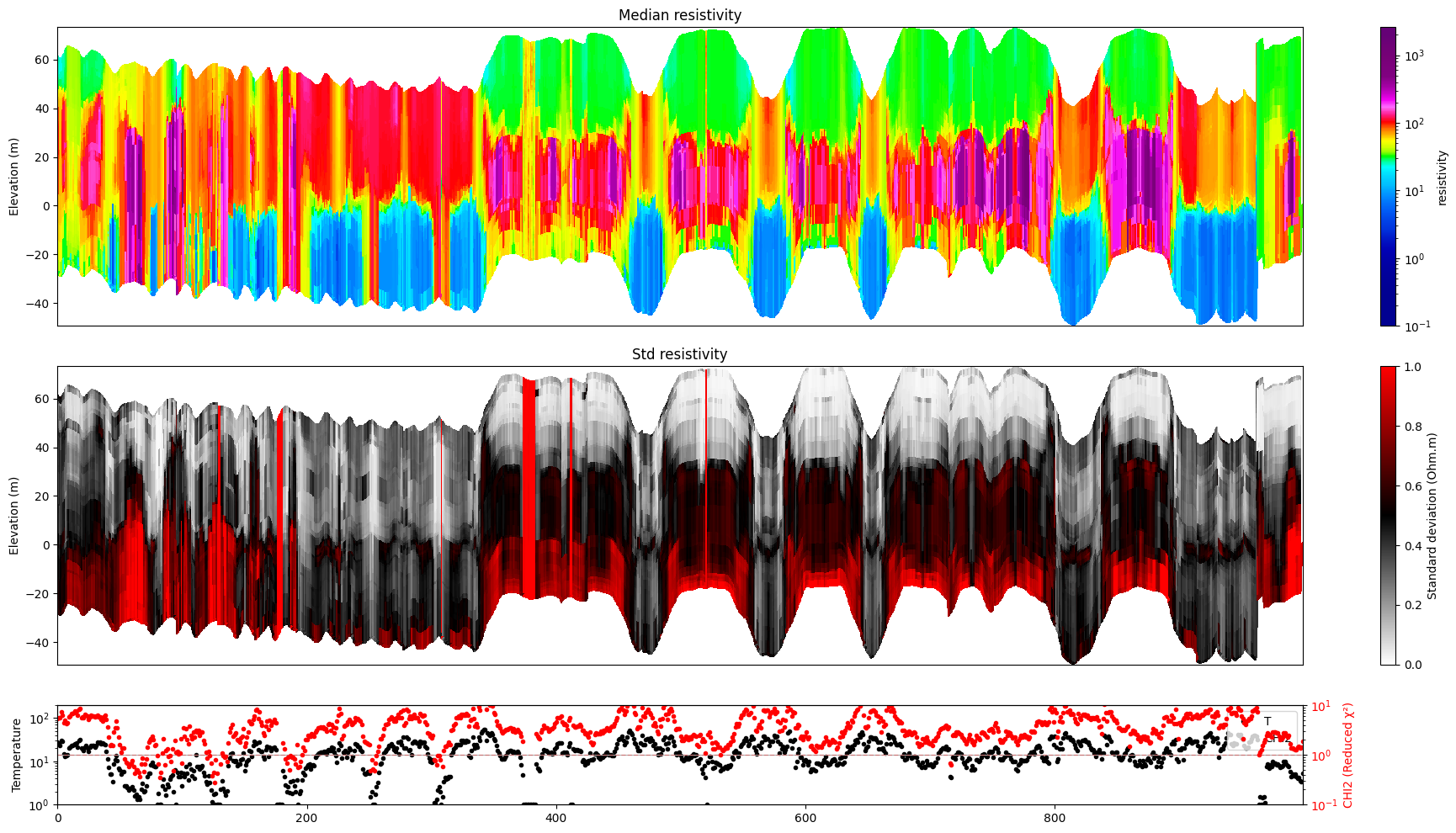
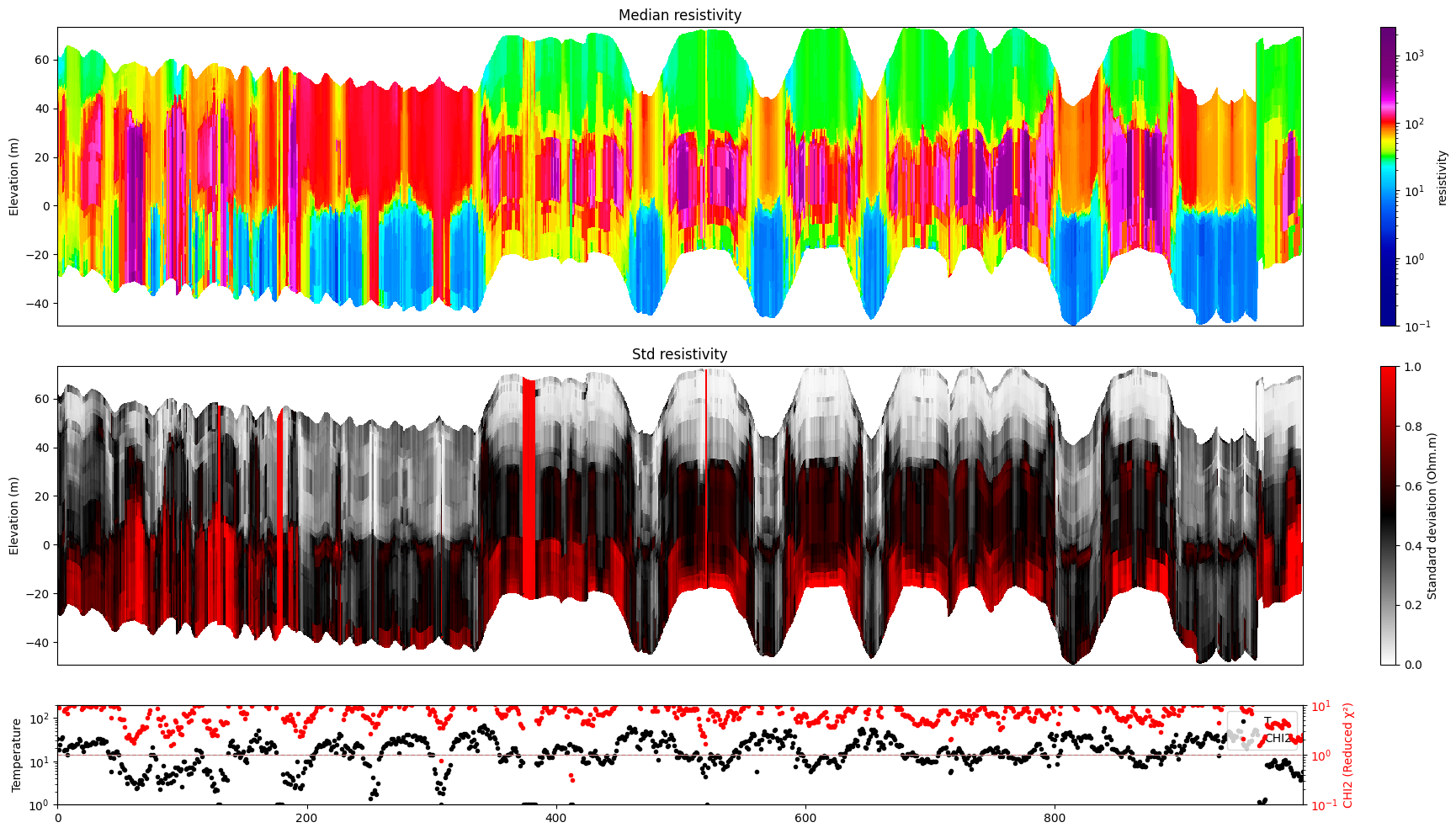
[13]:
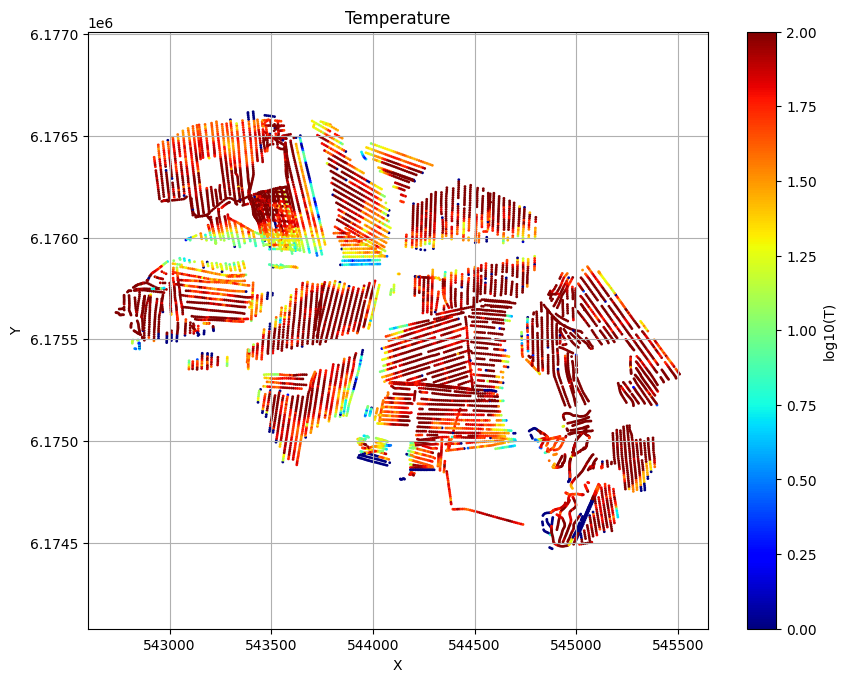
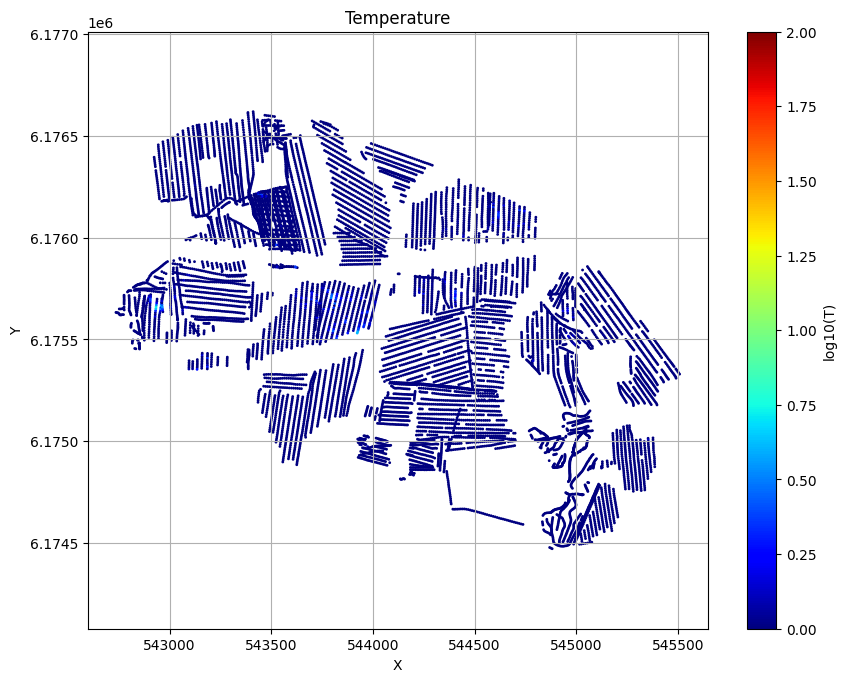
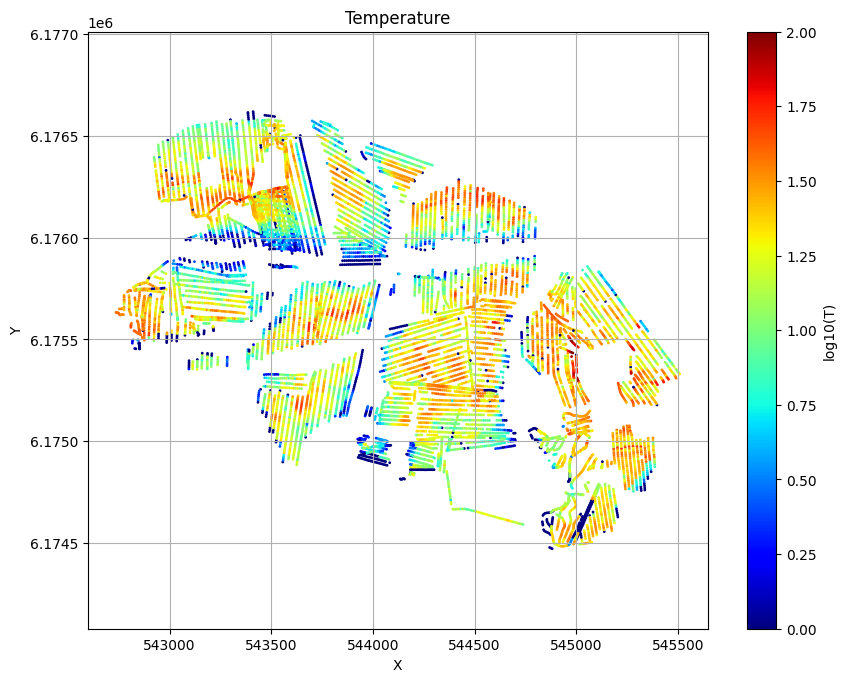
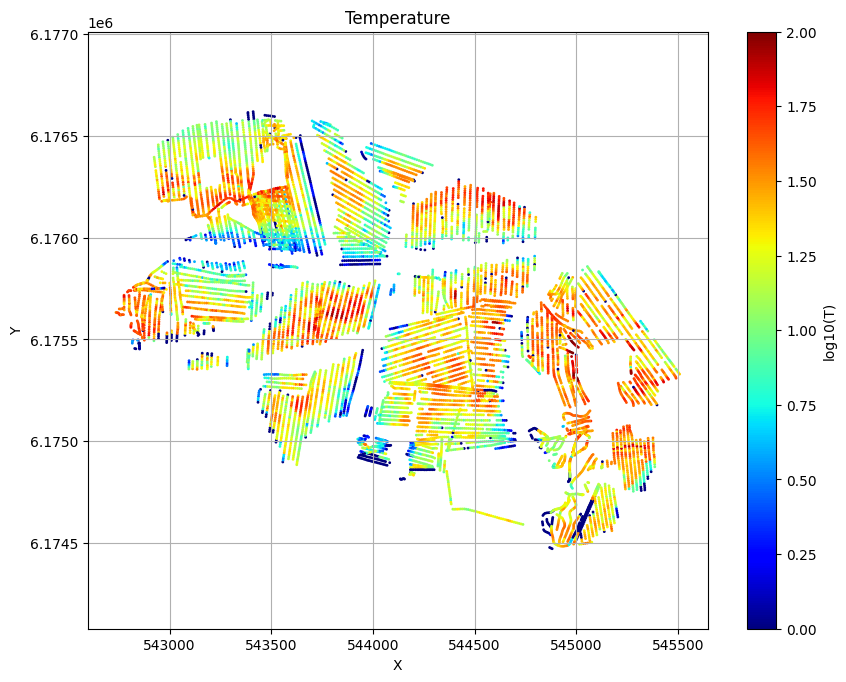
for f_post_h5 in f_post_arr:
ig.plot_T_EV(f_post_h5, pl='T', hardcopy=hardcopy)
plt.show()
#ig.plot_T_EV(f_post_h5, pl='EV', hardcopy=hardcopy)
[14]:
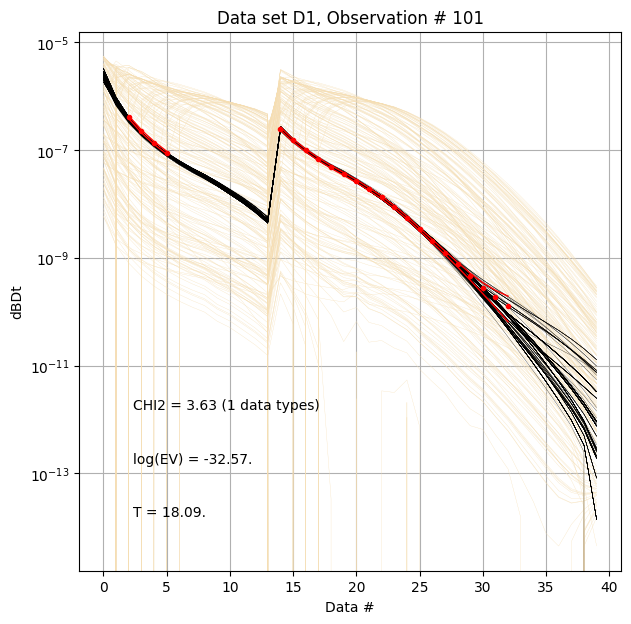
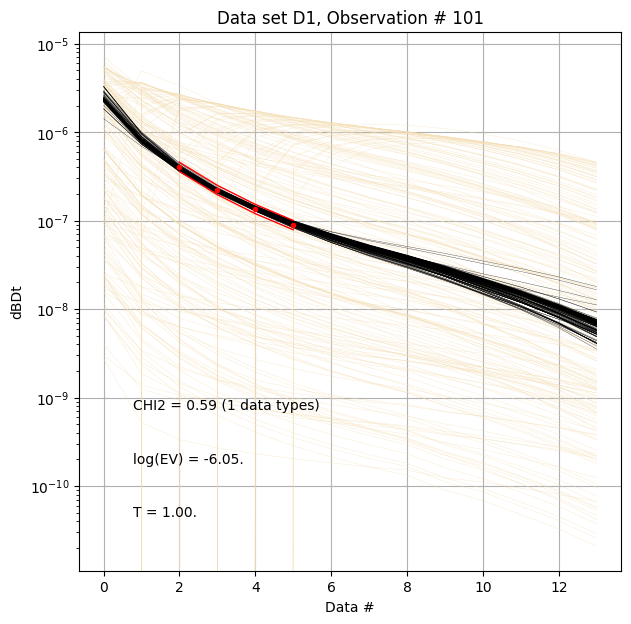
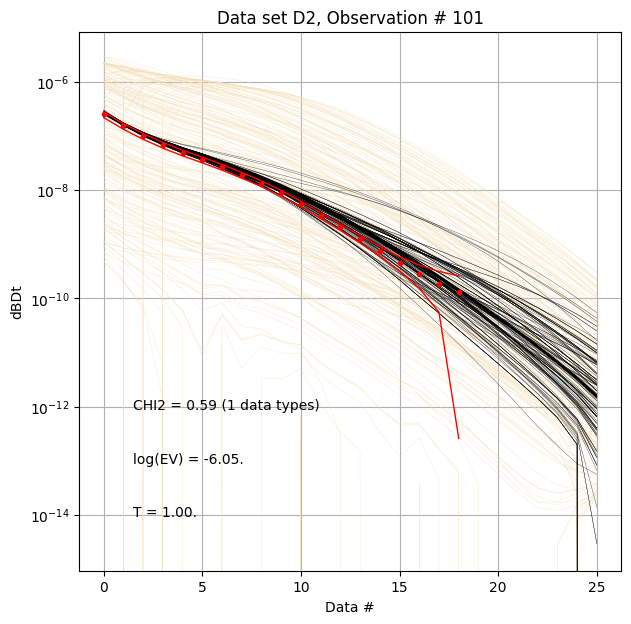
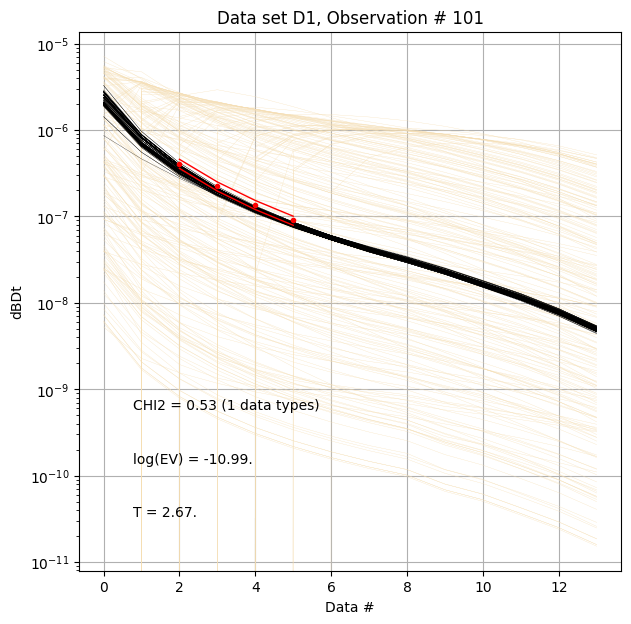
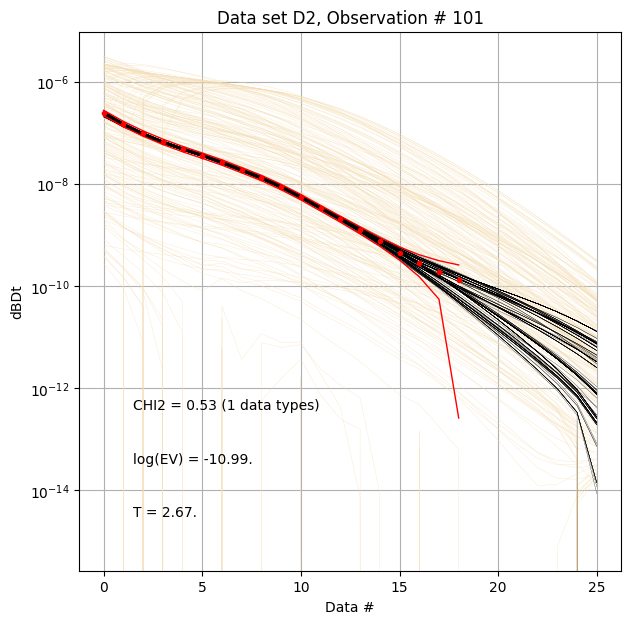
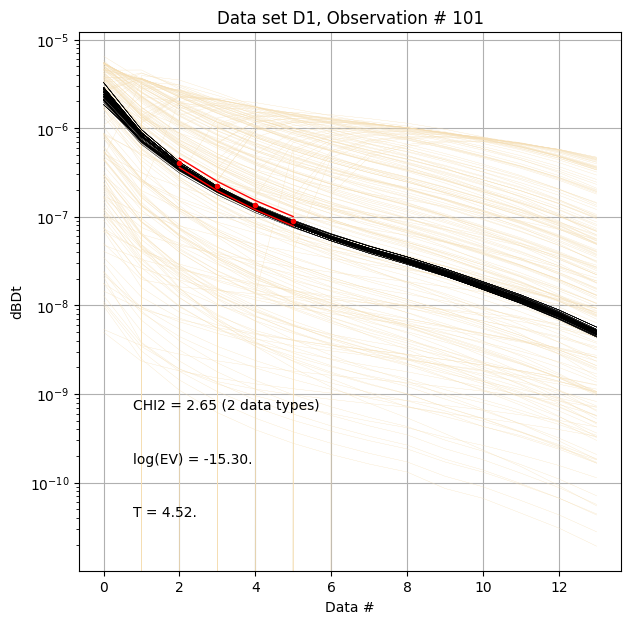
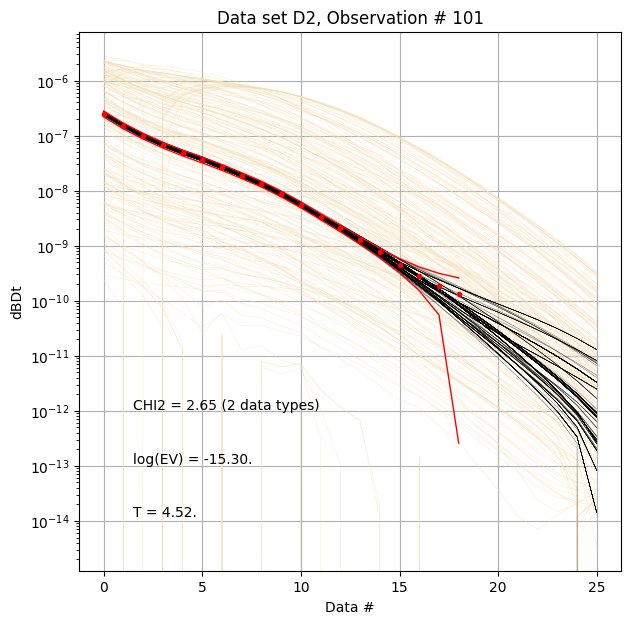
for f_post_h5 in f_post_arr:
# % Plot prior, posterior, and observed data
ig.plot_data_prior_post(f_post_h5, i_plot=100, hardcopy=hardcopy)
#ig.plot_data_prior_post(f_post_h5, i_plot=0, hardcopy=hardcopy)
[15]:
with h5py.File(f_data_h5, 'r') as f:
nobs = f['D1/d_obs'].shape[0]
ni=len(f_post_arr)
T = np.zeros((ni,nobs))
EV = np.zeros((ni,nobs))
EV_post = np.zeros((ni,nobs))
N_UNIQUE = np.zeros((ni,nobs))
i=-1
for f_post_h5 in f_post_arr:
with h5py.File(f_post_h5, 'r') as f:
i=i+1
T[i] = f['/T'][:]
EV[i] = f['/EV'][:]
N_UNIQUE[i] = f['/N_UNIQUE'][:]
EV_post[i] = f['/EV_post'][:]
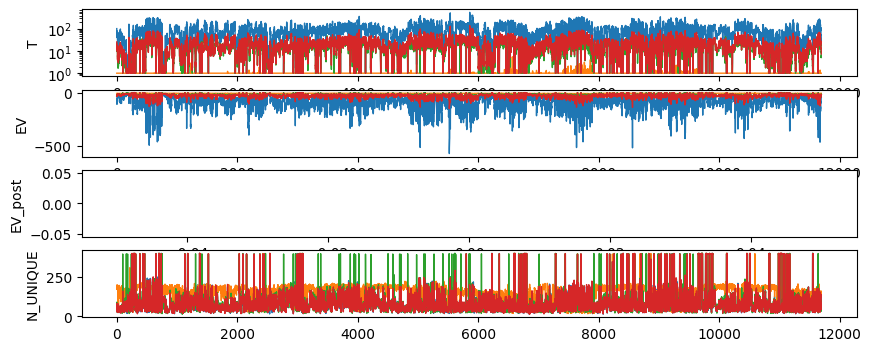
fig, ax = plt.subplots(4,1,figsize=(10,4))
ax[0].semilogy(T.T,'-', linewidth=1)
ax[0].set_ylabel('T')
ax[1].plot(EV.T,'-', linewidth=1)
ax[1].set_ylabel('EV')
ax[2].plot(EV_post.T,'-', linewidth=1)
ax[2].set_ylabel('EV_post')
ax[3].plot(N_UNIQUE.T,'-', linewidth=1)
ax[3].set_ylabel('N_UNIQUE')
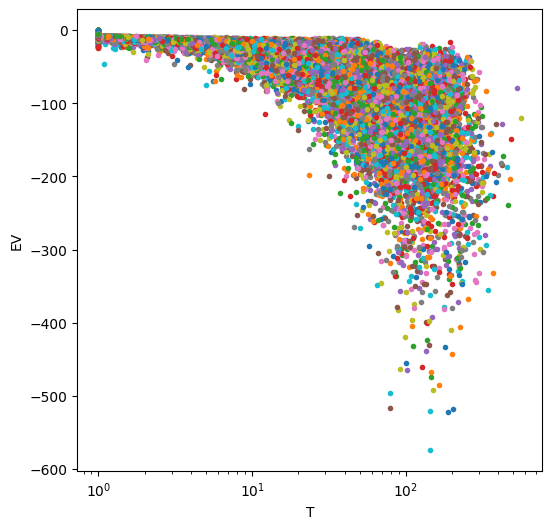
fig=plt.figure(figsize=(6,6))
plt.semilogx(T,EV,'.')
plt.xlabel('T')
plt.ylabel('EV')
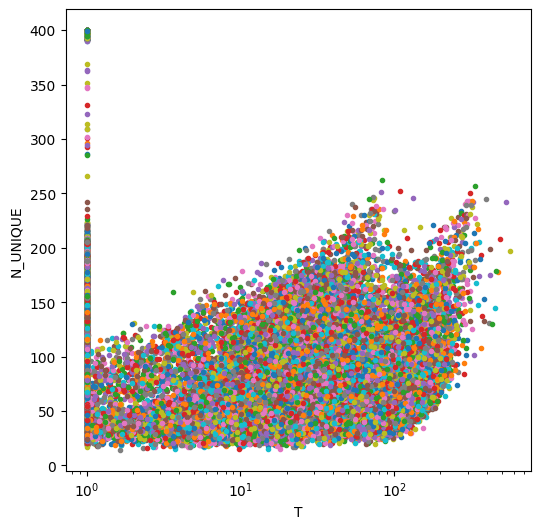
fig=plt.figure(figsize=(6,6))
plt.semilogx(T,N_UNIQUE,'.')
plt.xlabel('T')
plt.ylabel('N_UNIQUE')
[15]:
Text(0, 0.5, 'N_UNIQUE')
[16]:
#ig.post_to_csv(f_post_h5)