Merging multiple data files from the same survey (ESBJERG)¶
Data from Esbjerg was recorded in 8 individual surveys, using 4 different calibations (leading to 4 different system GEX files)
”TX07_20230906_2x4_RC20-33.gex”: 20230921, 20230922, 20230925, 20230926 “TX07_20231016_2x4_RC20-33.gex”: 20231026, 20231027 “TX07_20231127_2x4x1_RC20_33.gex”: 20240109 “TX07_20240125_2x4_RC20-33.gex”: 20240313
For the first 3 surveys the same number of gates were used, but it differedin the lasy (TX07_20240125_2x4_RC20-33.gex)
Below is an example of how to
A: invert each survey individually, and then combine the results
a1) combine survey files (data) for each calibration file
b1) invert each survey individually
c1) merge all posterior results in to one posterior file
B: Combine all data (for the first three surveys) and one system file to invert all data
[1]:
try:
# Check if the code is running in an IPython kernel (which includes Jupyter notebooks)
get_ipython()
# If the above line doesn't raise an error, it means we are in a Jupyter environment
# Execute the magic commands using IPython's run_line_magic function
get_ipython().run_line_magic('load_ext', 'autoreload')
get_ipython().run_line_magic('autoreload', '2')
except:
# If get_ipython() raises an error, we are not in a Jupyter environment
# # #%load_ext autoreload
# # #%autoreload 2
pass
[2]:
import integrate as ig
import numpy as np
import matplotlib.pyplot as plt
# check if parallel computations can be performed
parallel = ig.use_parallel(showInfo=1)
Notebook detected. Parallel processing is OK
GET The raw data, and merge surveys¶
[3]:
# 8 DATA files are available, corresponding to 8 different surveys obtained with 4 different GEX files
case = 'ESBJERG'
files = ig.get_case_data(case=case, loadType='premerge')
Getting data for case: ESBJERG
Downloading 20230921_AVG_export.h5
Downloaded 20230921_AVG_export.h5
Downloading 20230922_AVG_export.h5
Downloaded 20230922_AVG_export.h5
Downloading 20230925_AVG_export.h5
Downloaded 20230925_AVG_export.h5
Downloading 20230926_AVG_export.h5
Downloaded 20230926_AVG_export.h5
Downloading 20231026_AVG_export.h5
Downloaded 20231026_AVG_export.h5
Downloading 20231027_AVG_export.h5
Downloaded 20231027_AVG_export.h5
Downloading 20240109_AVG_export.h5
Downloaded 20240109_AVG_export.h5
Downloading 20240313_AVG_export.h5
Downloaded 20240313_AVG_export.h5
--> Got data for case: ESBJERG
Case A: Treat each area individually¶
[4]:
# Define the surveys with the same systemfile (gex)
f_gex1 ='TX07_20230906_2x4_RC20-33.gex'
f_data1 = []
f_data1.append('20230921_AVG_export.h5')
f_data1.append('20230922_AVG_export.h5')
f_data1.append('20230925_AVG_export.h5')
f_data1.append('20230926_AVG_export.h5')
f_gex2 ='TX07_20231016_2x4_RC20-33.gex'
f_data2 = []
f_data2.append('20231026_AVG_export.h5')
f_data2.append('20231027_AVG_export.h5')
f_gex3 = 'TX07_20231127_2x4x1_RC20_33.gex'
f_data3 = []
f_data3.append('20240109_AVG_export.h5')
f_gex4 = 'TX07_20240125_2x4_RC20-33.gex'
f_data4 = []
f_data4.append('20240313_AVG_export.h5')
[5]:
# Merge the data
f_data_merged1 = ig.merge_data(f_data1, f_gex1, f_data_merged_h5='ESBJERG_1.h5')
f_data_merged2 = ig.merge_data(f_data2, f_gex2, f_data_merged_h5='ESBJERG_2.h5')
f_data_merged3 = ig.merge_data(f_data3, f_gex3, f_data_merged_h5='ESBJERG_3.h5')
f_data_merged4 = ig.merge_data(f_data4, f_gex4, f_data_merged_h5='ESBJERG_4.h5')
Data has 20907 stations and 40 channels
Adding group ESBJERG_1.h5:D1
Data has 5208 stations and 40 channels
Adding group ESBJERG_2.h5:D1
Data has 1946 stations and 40 channels
Adding group ESBJERG_3.h5:D1
Data has 3693 stations and 39 channels
Adding group ESBJERG_4.h5:D1
[6]:
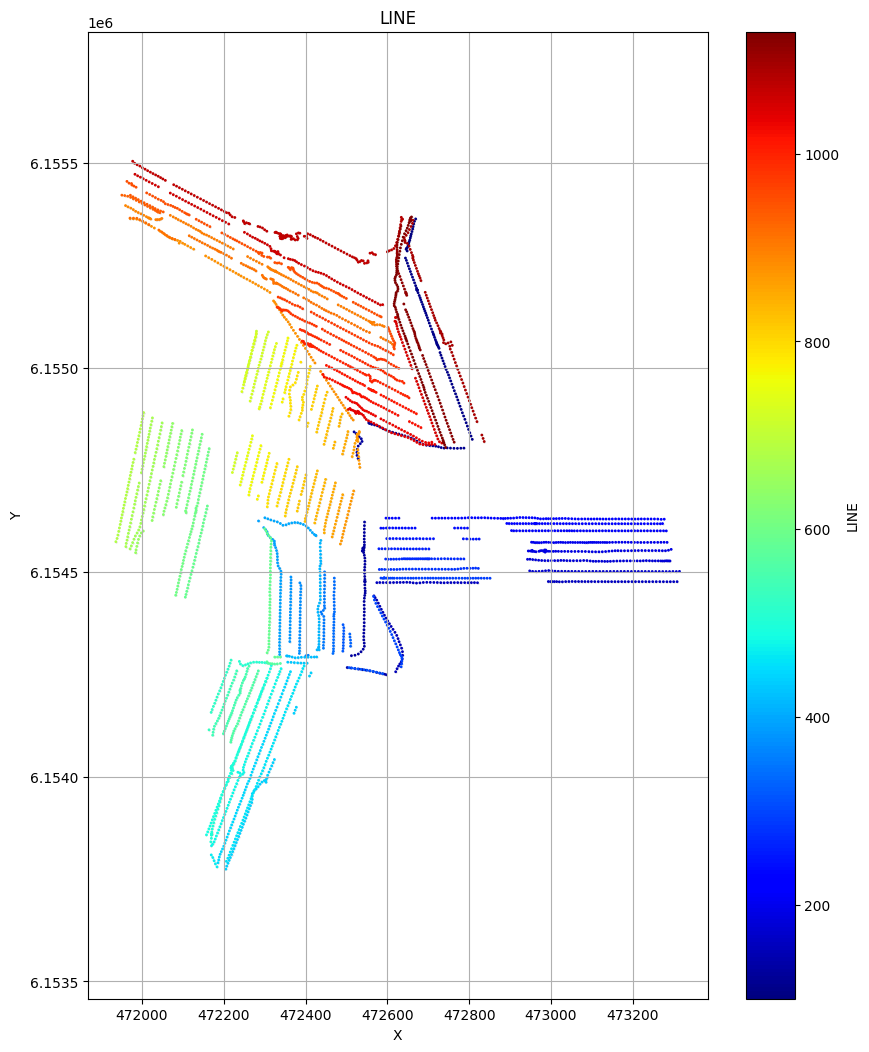
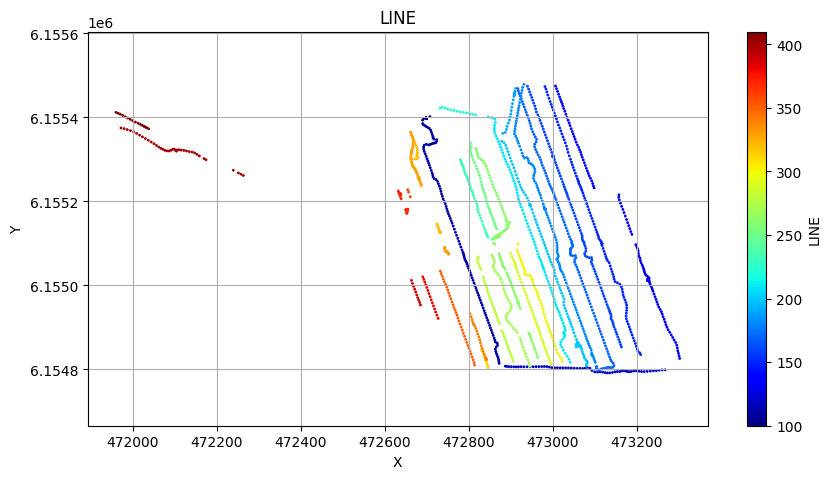
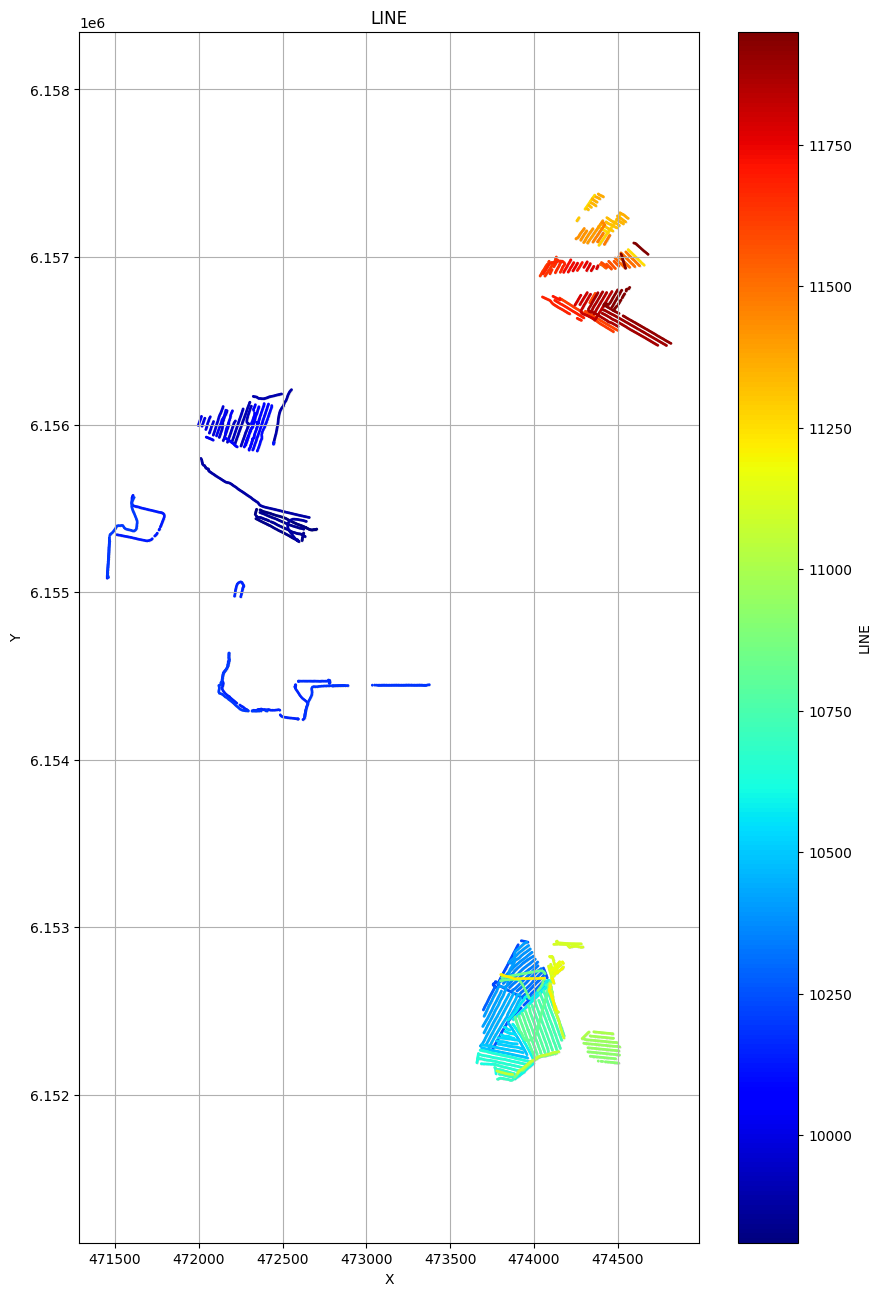
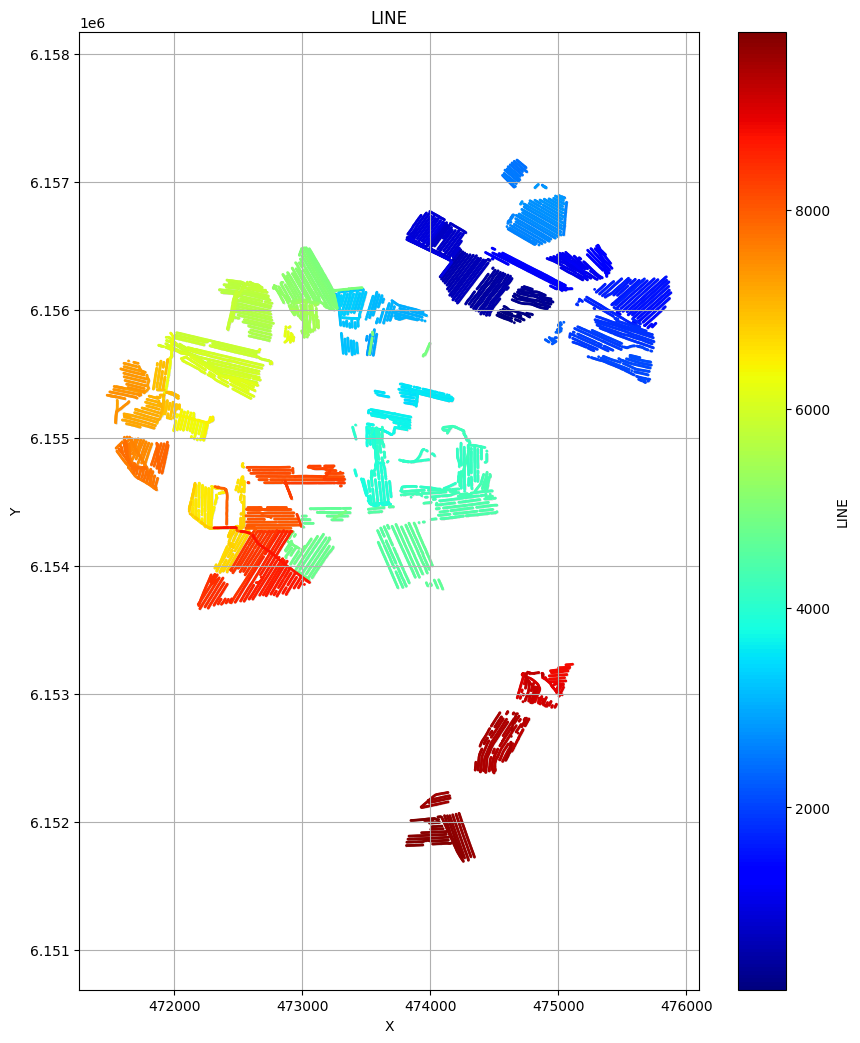
# Plot the geometry for each of the 4 areas
ig.plot_geometry(f_data_merged4, pl='LINE')
ig.plot_geometry(f_data_merged3, pl='LINE')
ig.plot_geometry(f_data_merged2, pl='LINE')
ig.plot_geometry(f_data_merged1, pl='LINE')
f_data_h5=ESBJERG_4.h5
f_data_h5=ESBJERG_3.h5
f_data_h5=ESBJERG_2.h5
f_data_h5=ESBJERG_1.h5
Run multiple inversions¶
[7]:
# Set PRIOR model: The prior is the same for all sets of data
N=100000
f_prior_h5 = ig.prior_model_layered(N=N,lay_dist='chi2', NLAY_deg=4, RHO_min=1, RHO_max=3000)
# Go through each of the 4 areas and invert the data seperately, using the same prior model, but different GEX files and hence prior data
f_data_all = [f_data_merged1, f_data_merged2, f_data_merged3, f_data_merged4]
f_prior_data_all=[]
f_post_all=[]
for f_data_h5 in f_data_all:
file_gex = ig.get_gex_file_from_data(f_data_h5)
f_prior_data_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex, parallel=parallel, showInfo=0)
#f_prior_data_h5 = ig.prior_data_gaaem(f_prior_h5, f_data_h5, parallel=parallel, showInfo=0)
f_prior_data_all.append(f_prior_data_h5)
f_post_h5 = ig.integrate_rejection(f_prior_data_h5,
f_data_h5,
showInfo=0,
parallel=parallel)
#updatePostStat=False)
f_post_all.append(f_post_h5)
ig.plot_feature_2d(f_post_h5,im=1,iz=20, key='Median', uselog=1, cmap='jet', s=1)
plt.show()
File PRIOR_CHI2_NF_4_log-uniform_N100000.h5 does not exist.
Using file_basename=TX07_20230906_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 90.2s/100000 soundings. 0.9ms/sounding, 1108.3it/s
integrate_rejection: Time=144.7s/20907 soundings, 6.9ms/sounding, 144.4it/s. T_av=53.2, EV_av=-74.6
/M1/Median[20,:] resistivity
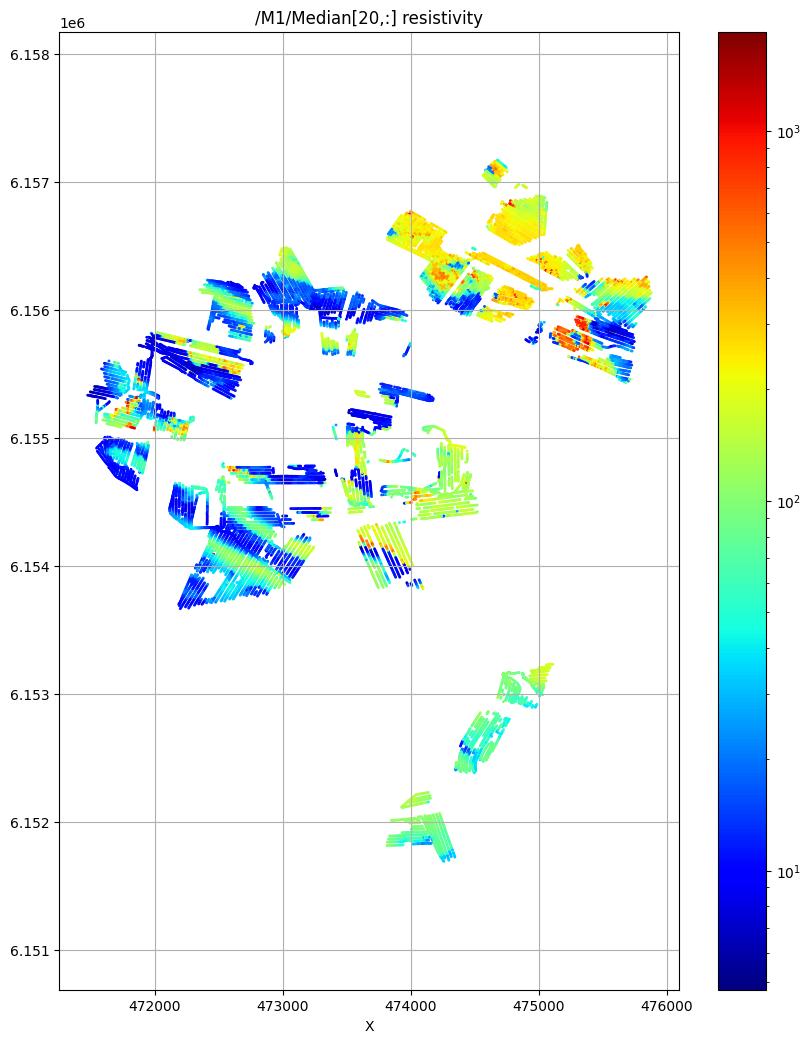
Using file_basename=TX07_20231016_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 89.0s/100000 soundings. 0.9ms/sounding, 1123.9it/s
integrate_rejection: Time= 36.2s/5208 soundings, 7.0ms/sounding, 143.7it/s. T_av=45.8, EV_av=-66.8
/M1/Median[20,:] resistivity
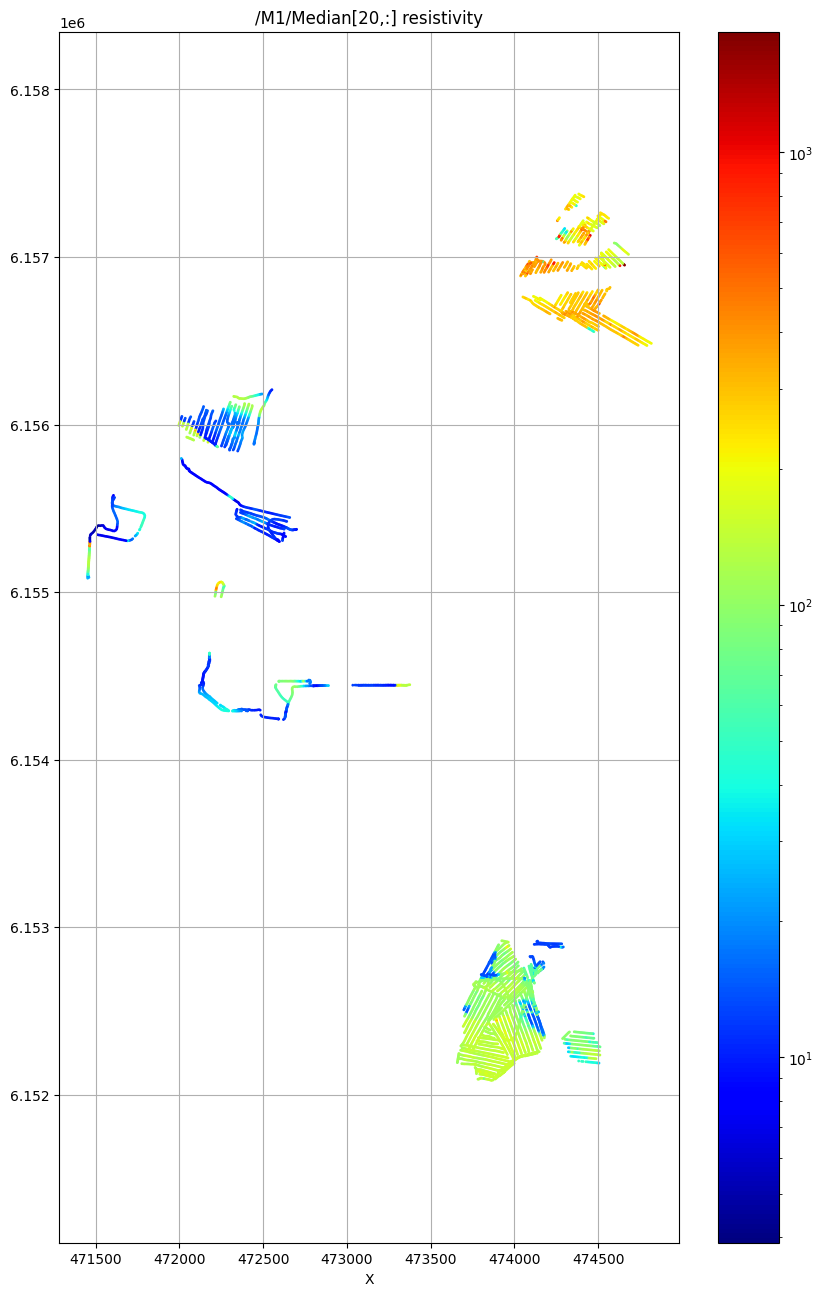
Using file_basename=TX07_20231127_2x4x1_RC20_33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 88.7s/100000 soundings. 0.9ms/sounding, 1128.0it/s
integrate_rejection: Time= 13.8s/1946 soundings, 7.1ms/sounding, 141.5it/s. T_av=18.1, EV_av=-25.9
/M1/Median[20,:] resistivity
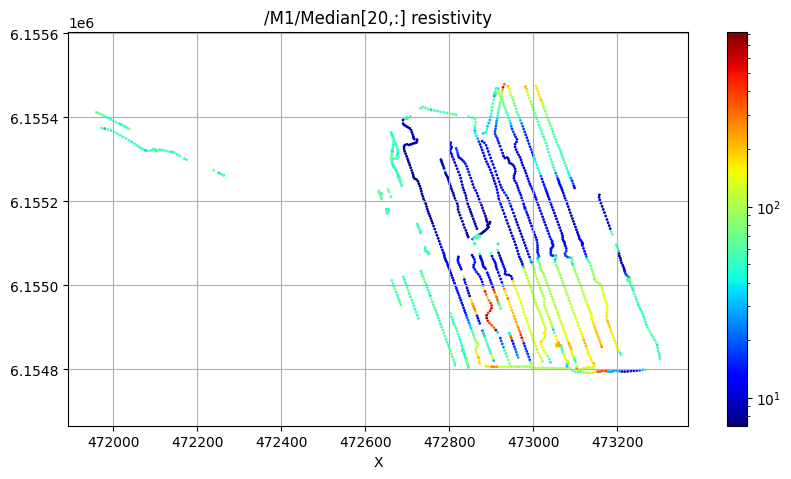
Using file_basename=TX07_20240125_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 89.6s/100000 soundings. 0.9ms/sounding, 1115.7it/s
integrate_rejection: Time= 24.0s/3693 soundings, 6.5ms/sounding, 153.9it/s. T_av=48.7, EV_av=-67.1
/M1/Median[20,:] resistivity

Merge the posterior data into one posterior¶
[8]:
f_post_merged_h5, f_data_merged_h5 = ig.merge_posterior(f_post_all[0:-2], f_data_all[0:-2], f_post_merged_h5='POST_ESBJERG_MERGED.h5')
ig.integrate_posterior_stats(f_post_merged_h5)
ig.plot_feature_2d(f_post_merged_h5,im=1,iz=20, key='Median', uselog=1, cmap='jet', s=1)
plt.show()
Data has 26115 stations and 40 channels
Adding group DATA_merged_N2.h5:D1
/M1/Median[20,:] resistivity
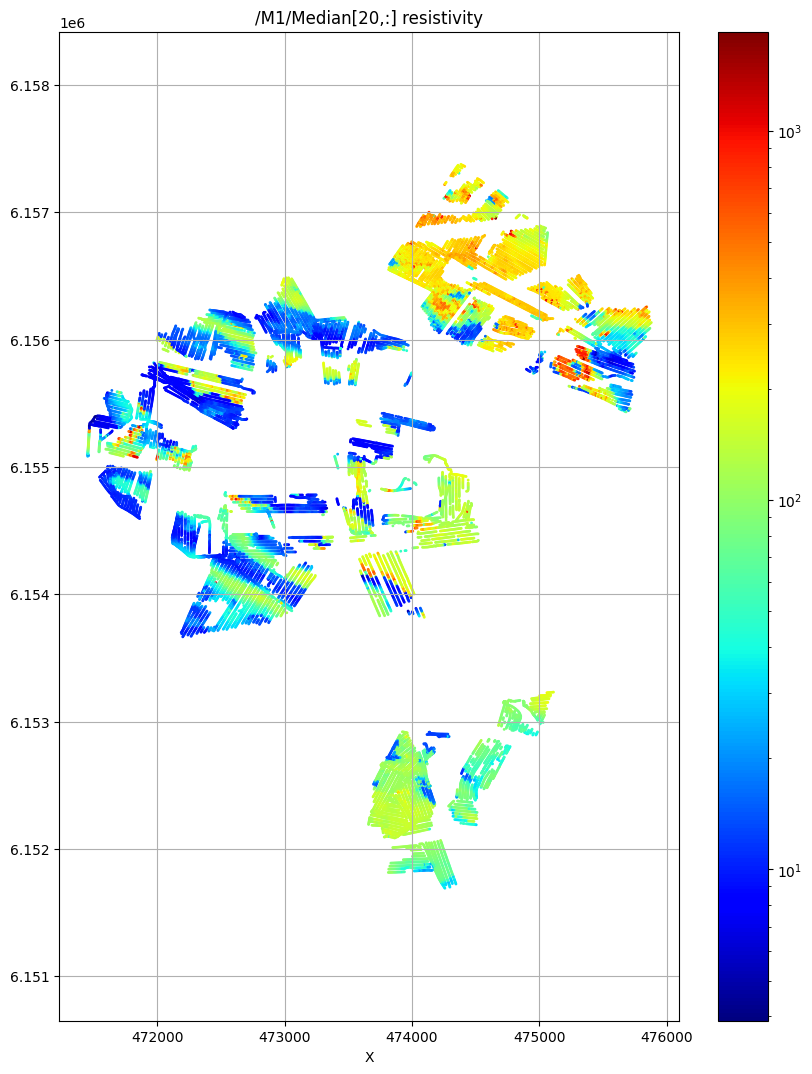
Case B: Combine all data, and use a single system GEX file¶
[9]:
# Combine f_data1, f_data2, and f_data3 into a single data set, and use a single system GEX file (this leads to an error)
## Cannot used the last dataset witjh rest as it has a different form!!!
# f_data = f_data1 + f_data2 + f_data3 + f_data4
f_data = f_data1 + f_data2 + f_data3
f_data_all_h5 = ig.merge_data(f_data1+f_data2+f_data3, f_gex1, f_data_merged_h5='ESBJERG_ALL.h5')
#ig.plot_geometry(f_data_merged_h5, pl='LINE')
Data has 28061 stations and 40 channels
Adding group ESBJERG_ALL.h5:D1
[10]:
file_gex = ig.get_gex_file_from_data(f_data_all_h5)
f_prior_data_all_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex, parallel=parallel, showInfo=0)
f_post_all_h5 = ig.integrate_rejection(f_prior_data_all_h5,
f_data_all_h5,
showInfo=0,
parallel=parallel,
updatePostStat=True)
Using file_basename=TX07_20230906_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time= 88.9s/100000 soundings. 0.9ms/sounding, 1125.3it/s
integrate_rejection: Time=194.6s/28061 soundings, 6.9ms/sounding, 144.2it/s. T_av=49.4, EV_av=-69.8
[11]:
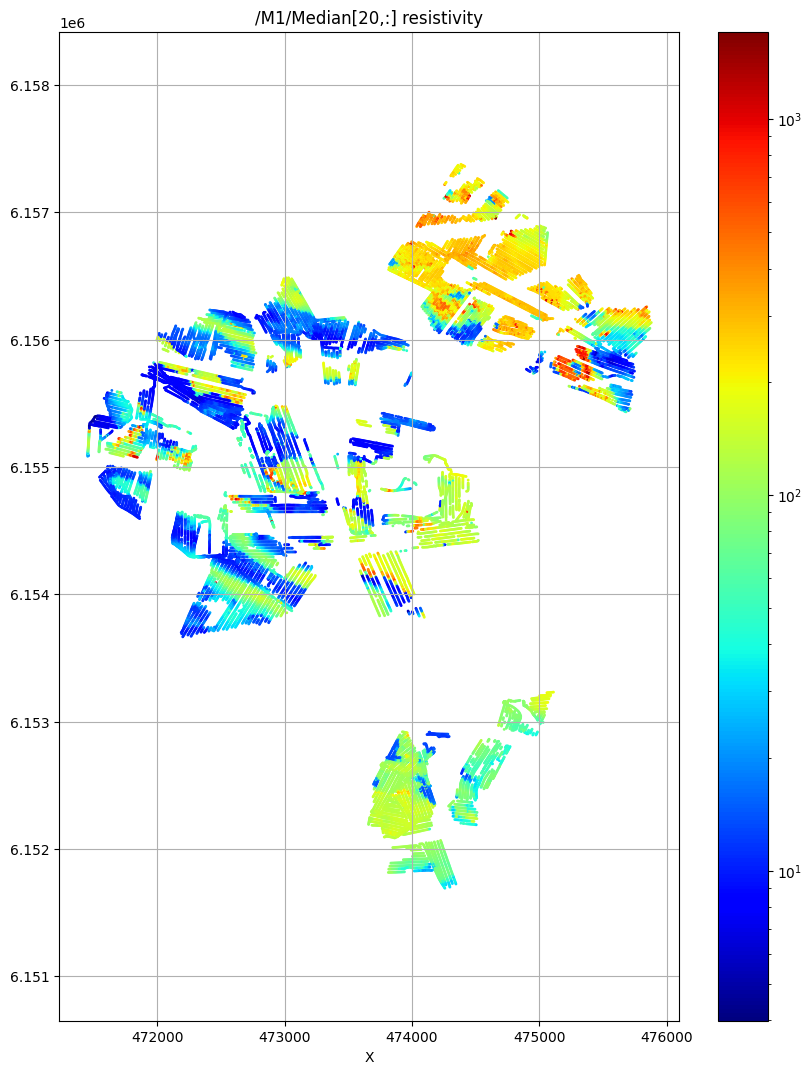
ig.plot_feature_2d(f_post_all_h5,im=1,iz=20, key='Median', uselog=1, cmap='jet', s=1)
plt.show()
/M1/Median[20,:] resistivity
[ ]:
[12]:
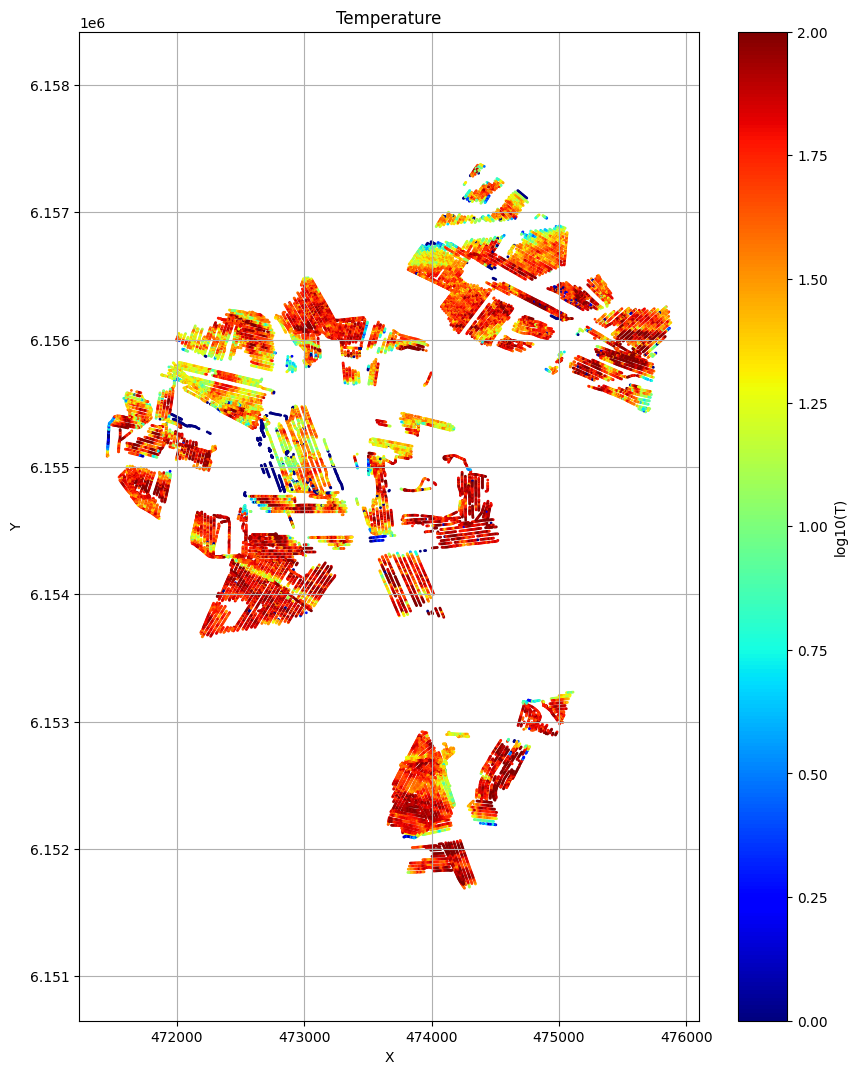
ig.plot_T_EV(f_post_all_h5, pl='T')
plt.show()
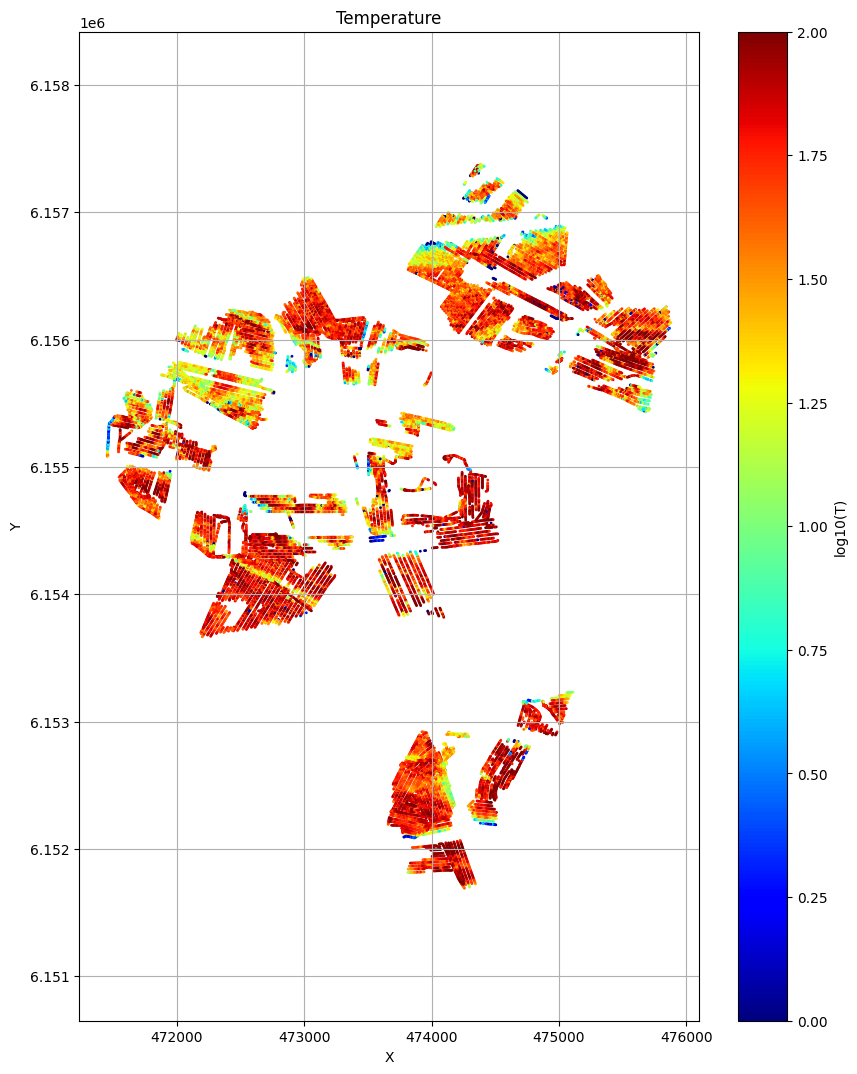
ig.plot_T_EV(f_post_merged_h5, pl='T')
plt.show()
[ ]: