Synthetic Case Study example with different noise models (uncorrelated/correlated)¶
Demonstrates the use of different noise models with tTEM data using an example using inverting data obtained from synthetic reference model
[1]:
try:
# Check if the code is running in an IPython kernel (which includes Jupyter notebooks)
get_ipython()
# If the above line doesn't raise an error, it means we are in a Jupyter environment
# Execute the magic commands using IPython's run_line_magic function
get_ipython().run_line_magic('load_ext', 'autoreload')
get_ipython().run_line_magic('autoreload', '2')
except:
# If get_ipython() raises an error, we are not in a Jupyter environment
# # # # # # # # # # # # # #%load_ext autoreload
# # # # # # # # # # # # # #%autoreload 2
pass
import integrate as ig
# check if parallel computations can be performed
parallel = ig.use_parallel(showInfo=1)
import numpy as np
import os
import matplotlib.pyplot as plt
import h5py
hardcopy=True
Notebook detected. Parallel processing is OK
Create The reference model and data¶
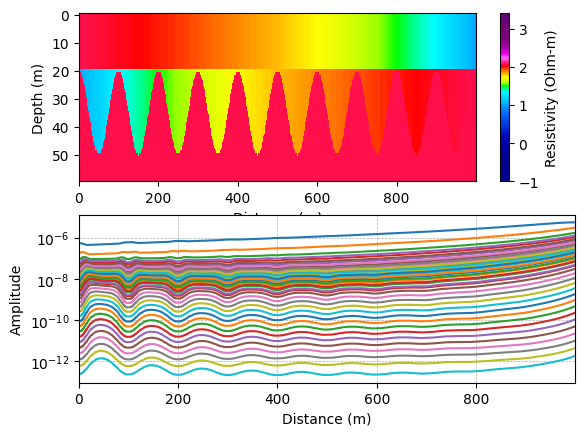
A simple three layer reference model is constructed, where the thickness of layer 2 varies. Them The corresponding noise free data is computed.
[2]:
# select the type of referenc model
case = '3layer' # A 3 layer referennce model
#case = 'wedge' # A 'wedge' reference model
z_max = 60 # Max thickness of the model
dx=1 # The layer thickness
rho = [120,10,120]
if case.lower() == 'wedge':
# Make Wedge MODEL
M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='Wedge', wedge_angle=10, z_max=z_max, dz=.5, x_max=100, dx=dx, z1=15, rho = rho)
M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='Wedge', wedge_angle=10, z_max=z_max, dz=.5, x_max=200, dx=dx, z1=15, rho = rho)
elif case.lower() == '3layer':
# Make 3 layer MODEL
M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='3layer', dx=dx, rho1_1 = rho[0], rho1_2 = rho[1], x_max = 100, x_range = 10)
#M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='3layer', dx=dx, rho1_1 = rho[0], rho1_2 = rho[1], x_max = 200, x_range = 20)
M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='3layer', dx=dx, rho1_1 = rho[0], rho1_2 = rho[1], x_max = 1000, x_range = 100)
M_ref, x_ref, z_ref, M_ref_lith, layer_depths = ig.synthetic_case(case='3layer', dx=dx, rho1_1 = rho[0], rho1_2 = rho[1], x_max = 1000, x_range = 50)
# Create reference data
f_data_h5 = '%s_%d' % (case,z_max)
thickness = np.diff(z_ref)
# Get a GEX file to use for creation of data
file_gex = ig.get_case_data(case='DAUGAARD', filelist=['TX07_20231016_2x4_RC20-33.gex'])[0]
# Compute the noise free reference data
D_ref = ig.forward_gaaem(C=1./M_ref, thickness=thickness, file_gex=file_gex)
Getting data for case: DAUGAARD
--> Got data for case: DAUGAARD
[3]:
M_ref.shape
[3]:
(1000, 60)
Plot the reference model and data¶
[4]:
cmap, clim = ig.get_colormap_and_limits(cmap_type='resistivity')
plt.subplot(2,1,1)
xx_ref, zz_ref = np.meshgrid(x_ref, z_ref)
#plt.pcolormesh(xx_ref, zz_ref, M_ref.T, cmap='jet', vmin=clim[0], vmax=clim[1])
plt.pcolormesh(xx_ref, zz_ref, np.log10(M_ref.T), cmap=cmap, vmin=np.log10(clim[0]), vmax=np.log10(clim[1]))
plt.xlim([x_ref.min(), x_ref.max()])
plt.xlabel('Distance (m)')
plt.ylabel('Depth (m)')
#plt.axis('equal')
plt.gca().invert_yaxis()
plt.colorbar(label='Resistivity (Ohm-m)')
plt.subplot(2,1,2)
plt.semilogy(x_ref, D_ref)
plt.xlim([x_ref.min(), x_ref.max()])
plt.xlabel('Distance (m)')
plt.ylabel('Amplitude')
plt.grid(True, which='both', linestyle='--', linewidth=0.5)
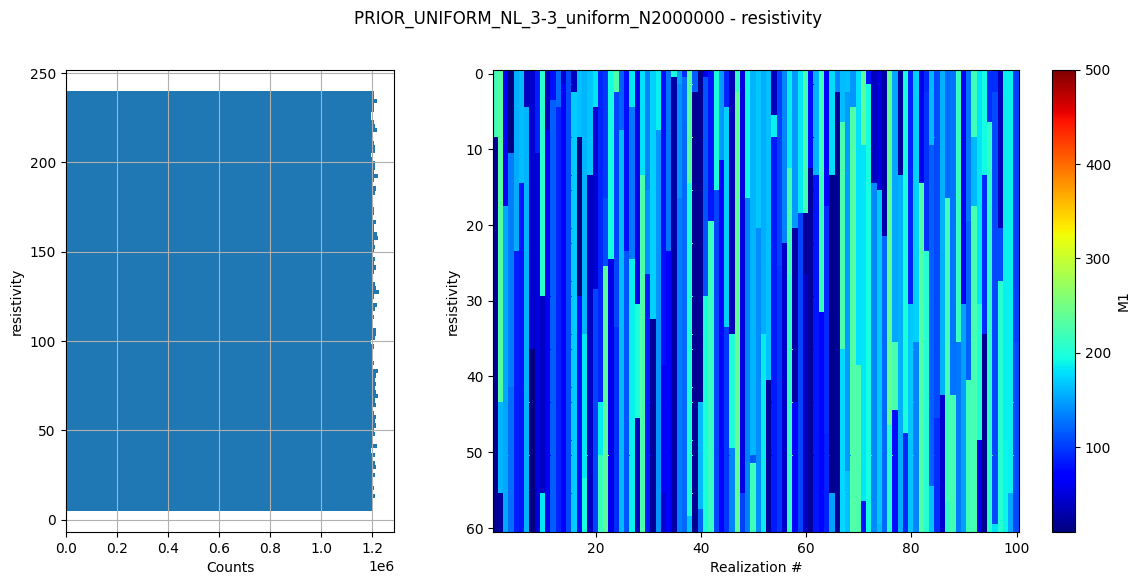
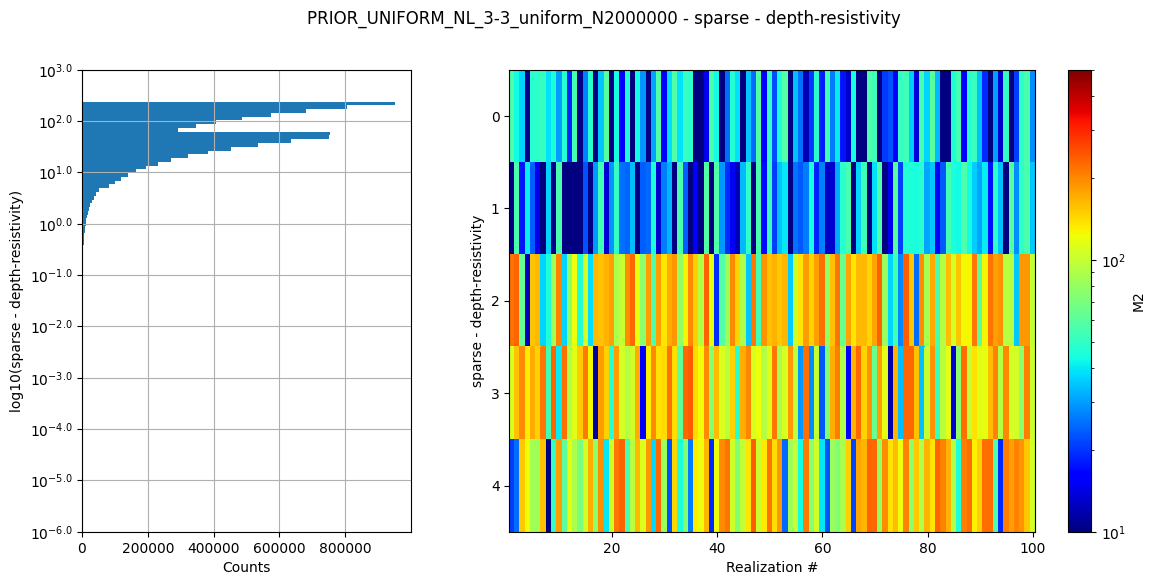
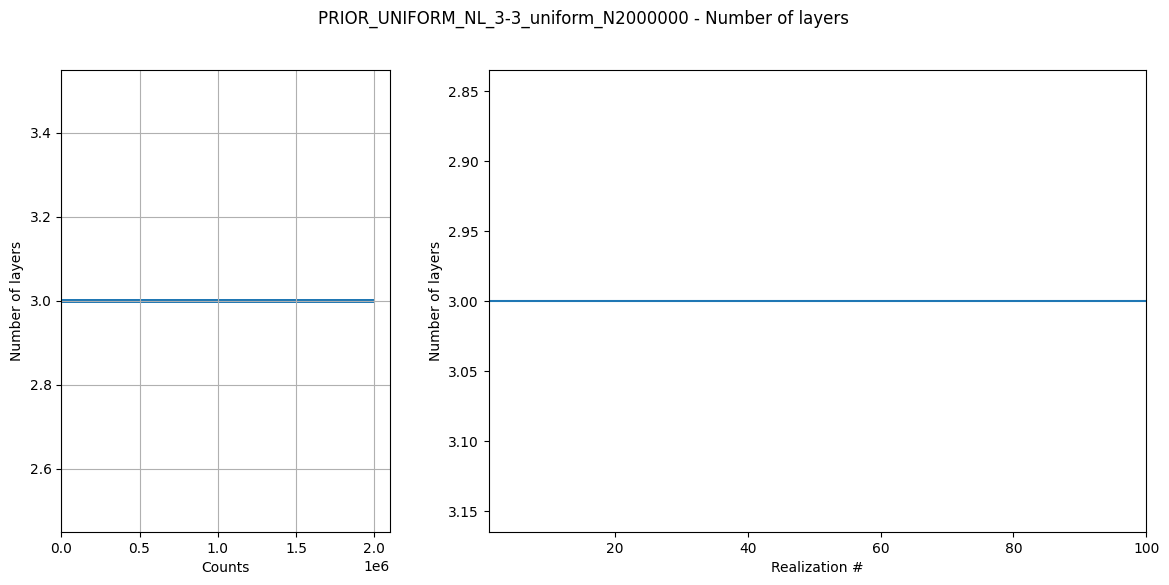
Create prior model and data¶
Here a prior model is defined (a 3 layer model). The prior model realizations are generated, followed by prior data realizations.
[5]:
N=2000000 # sample size
#N= 100000 # sample size
NLAY_min=3
NLAY_max=3
f_prior_data_h5='PRIOR_UNIFORM_NL_%d-%d_uniform_N%d_TX07_20231016_2x4_RC20-33_Nh280_Nf12.h5' % (NLAY_min, NLAY_max, N)
# make prior model realizations
f_prior_h5 = ig.prior_model_layered(N=N,
lay_dist='uniform', z_max = z_max,
NLAY_min=NLAY_min, NLAY_max=NLAY_max,
RHO_dist='uniform', RHO_min=0.5*min(rho), RHO_max=2*max(rho))
# make prior data realizations
f_prior_data_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex)
# Plot some statistics about the prior model parameters
ig.plot_prior_stats(f_prior_h5)
Using file_basename=TX07_20231016_2x4_RC20-33
prior_data_gaaem: Using 32 parallel threads.
prior_data_gaaem: Time=2876.1s/2000000 soundings. 1.4ms/sounding, 695.4it/s
Three ways to describe uncorrelated noise¶
[6]:
# The basic noise model: 3% relative noise and a noise floor at 1e-12
rng = np.random.default_rng()
d_std = 0.03 # standard deviation of the noise
d_std_base = 1e-12 # base noise
D_std = d_std * D_ref + d_std_base
# Add a realization of of the noise model to the noise free data, and treat these as 'observed' reference data.
D_noise = rng.normal(0, D_std, D_ref.shape)
D_obs = D_ref + D_noise
# Cd is a diagnoal matrix with the standard deviation of the data
Setup data with different types of noise assumptions.¶
[7]:
# If a single correlated noise model is used,
# it can represented by be the mean of the standard deviation of the data.
# This is though an approximation.
Cd_single = np.diag(np.mean(D_std, axis=0)**2)
Cd_single = np.diag(D_std[0]**2)
# The full data covariance matrix is represented by a 3D array of shape (ns,nd,nd)
# Using this type of noise should provide identical results to using d_std, only slower as
# the full covariance matrix is used.
# This type of noise model is useful when the noise is not the same for all data points,
# and the noise is correlated.
ns,nd=D_std.shape
Cd_mul = np.zeros((ns,nd,nd))
for i in range(ns):
Cd_mul[i] = np.diag(D_std[i]**2)
# Wrie the three differet types of noise models to hdf5 files
f_data_h5_arr=[]
name_arr = []
f_out = ig.save_data_gaussian(D_obs, D_std = D_std, f_data_h5 = 'data_uncorr.h5', id=1, showInfo=0, delete_if_exist=True)
f_data_h5_arr.append(f_out)
name_arr.append('Uncorrelated noise')
f_out = ig.save_data_gaussian(D_obs, Cd=Cd_single, f_data_h5 = 'data_corr1.h5', id=1, showInfo=0, delete_if_exist=True)
f_data_h5_arr.append(f_out)
name_arr.append('Correlated noise - mean')
f_out = ig.save_data_gaussian(D_obs, Cd=Cd_mul, f_data_h5 = 'data_corr2.h5', id=1, showInfo=0, delete_if_exist=True)
f_data_h5_arr.append(f_out)
name_arr.append('Correlated noise - individual')
Data has 1000 stations and 40 channels
Adding group data_uncorr.h5:D1
Data has 1000 stations and 40 channels
Adding group data_corr1.h5:D1
Data has 1000 stations and 40 channels
Adding group data_corr2.h5:D1
[8]:
import time as time
# test likelhood
doTest = True
if doTest:
id=0
d_obs = D_obs[id]
#d_obs[11]=np.nan
d_std = D_std[id]
C_single = np.diag(d_std**2)+1e-18
with h5py.File(f_prior_data_h5, 'r') as f:
D = f['/D1'][:]
#D = D_ref
t0=time.time()
L1 = ig.likelihood_gaussian_diagonal(D, d_obs, d_std)
t1 = time.time()-t0
L2 = ig.likelihood_gaussian_full(D, d_obs, Cd_single, useVectorized=True)
t2 = time.time()-t0-t1
#L3 = ig.likelihood_gaussian_full(D, d_obs, Cd_single, N_app = 110, checkNaN=True, useVectorized=False)
#L3 = ig.likelihood_gaussian_full(D, d_obs, Cd_mul[id], N_app = 1000, useVectorized=True)
L3 = ig.likelihood_gaussian_full(D, d_obs, Cd_mul[id], useVectorized=True)
t3 = time.time()-t2-t1-t0
# L3 is a list of log-likelihood values. I would like to compute the log of the mean of the likelihood values, that is
# the log(mean(exp(L3))). exp(L3) may lead to such small numbers that this becomes NaN. Using log-sum-exp trick for numerical stability.
def log_mean_exp(log_vals):
"""Compute log(mean(exp(log_vals))) using the log-sum-exp trick for numerical stability"""
max_val = np.max(log_vals)
return max_val + np.log(np.mean(np.exp(log_vals - max_val)))
mean_L1 = log_mean_exp(L1)
mean_L2 = log_mean_exp(L2)
mean_L3 = log_mean_exp(L3)
mean_L1 = np.log(np.mean(np.exp(L1)))
#print("L1: %f, L2: %f, L3: %f" % (mean_L1, mean_L2, mean_L3))
#¤
#
#print("L1: %f, L2: %f, L3: %f" % (L1[0], L2[0], L3[0]))
print("mean T1=%3.5f" % (mean_L1))
print("mean T2=%3.5f" % (mean_L2))
print("mean T3=%3.5f" % (mean_L3))
print("t1, t2, t3 = %f, %f, %f" % (t1, t2, t3))
print("SLOWDOWN = %f, %f, %f" % (t1/t1, t2/t1, t3/t1))
plt.semilogy(-L1, 'k.', label='L1', markersize=10)
plt.plot(-L2, 'b.', label='L2', markersize=5)
plt.plot(-L3, 'r.', label='L3', markersize=2)
plt.legend()
plt.ylabel('-log(L)')
plt.show()
mean T1=-40.42736
mean T2=-40.42736
mean T3=-40.42736
t1, t2, t3 = 1.039125, 2.156658, 2.157610
SLOWDOWN = 1.000000, 2.075456, 2.076372
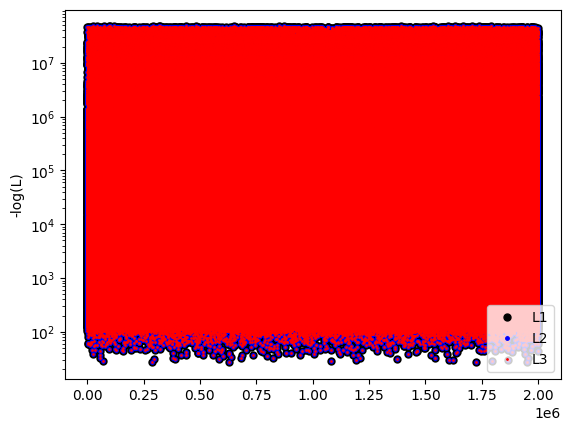
Invert each data set with different way to represent the noise.¶
[9]:
import time as time
f_post_h5_arr = []
T_arr = []
EV_arr = []
EV_post_arr = []
clim = [min(rho)*0.8, max(rho)*1.25]
t_elapsed = []
for f_data_h5 in f_data_h5_arr:
t0 = time.time()
f_post_h5 = ig.integrate_rejection(f_prior_data_h5, f_data_h5,
parallel=parallel,
Ncpu = 8,
#use_N_best=500
)
t_elapsed.append(time.time()-t0)
with h5py.File(f_post_h5, 'r') as f_post:
T_arr.append(f_post['/T'][:])
EV_arr.append(f_post['/EV'][:])
EV_post_arr.append(f_post['/EV_post'][:])
f_post_h5_arr.append(f_post_h5)
for i in range(len(name_arr)):
print('%s: t_elapsed = %f s' % (name_arr[i], t_elapsed[i]))
integrate_rejection: Time=296.4s/1000 soundings, 296.4ms/sounding, 3.4it/s. T_av=4.1, EV_av=-33.9
integrate_rejection: Time=494.1s/1000 soundings, 494.1ms/sounding, 2.0it/s. T_av=25.7, EV_av=-389.8
integrate_rejection: Time=497.7s/1000 soundings, 497.7ms/sounding, 2.0it/s. T_av=4.1, EV_av=-33.9
Uncorrelated noise: t_elapsed = 300.701822 s
Correlated noise - mean: t_elapsed = 498.756250 s
Correlated noise - individual: t_elapsed = 502.262018 s
[ ]:
[10]:
for i in range(len(f_post_h5_arr)):
ig.plot_profile(f_post_h5_arr[i],hardcopy=hardcopy, clim = clim, im=1)
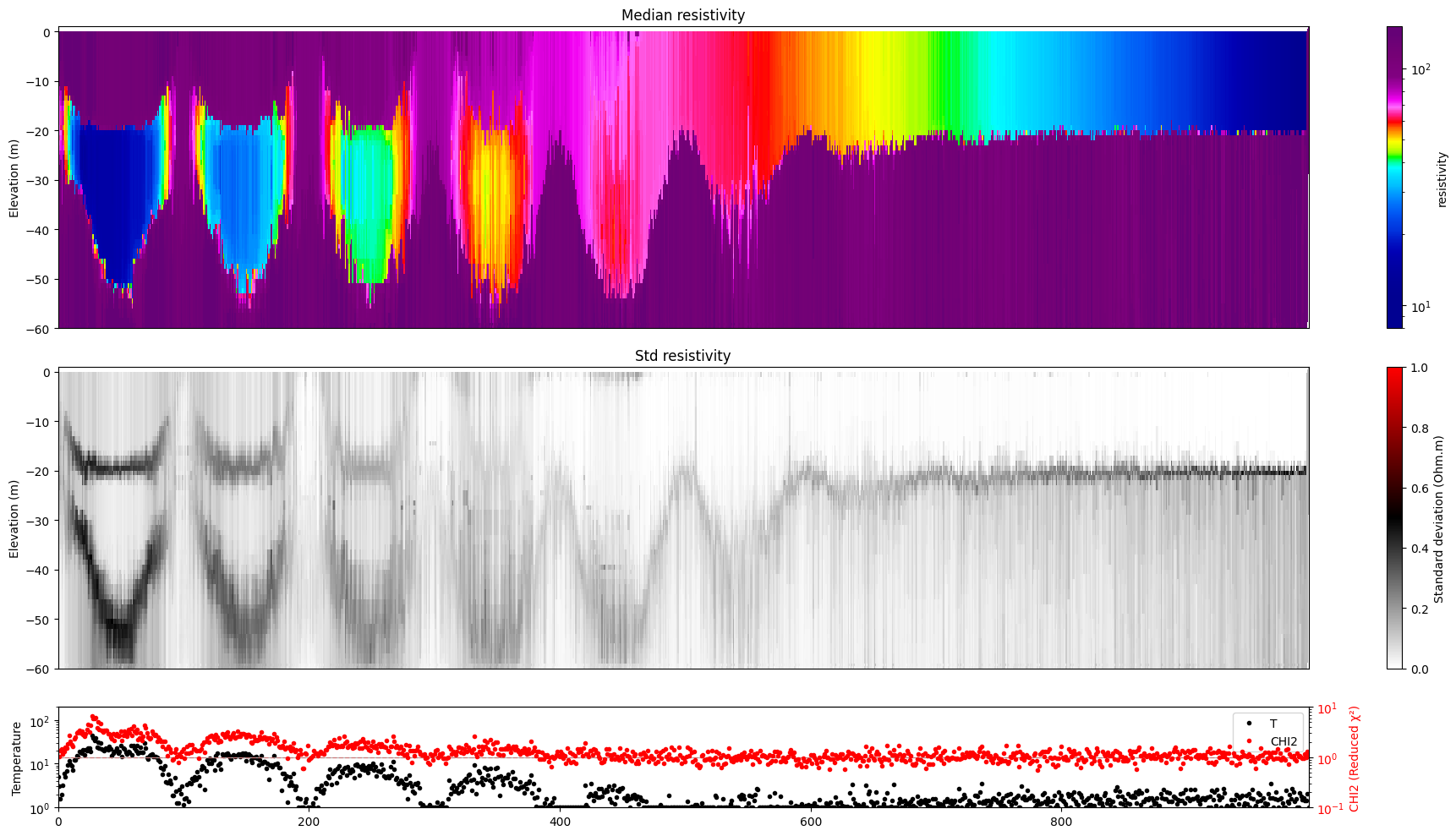
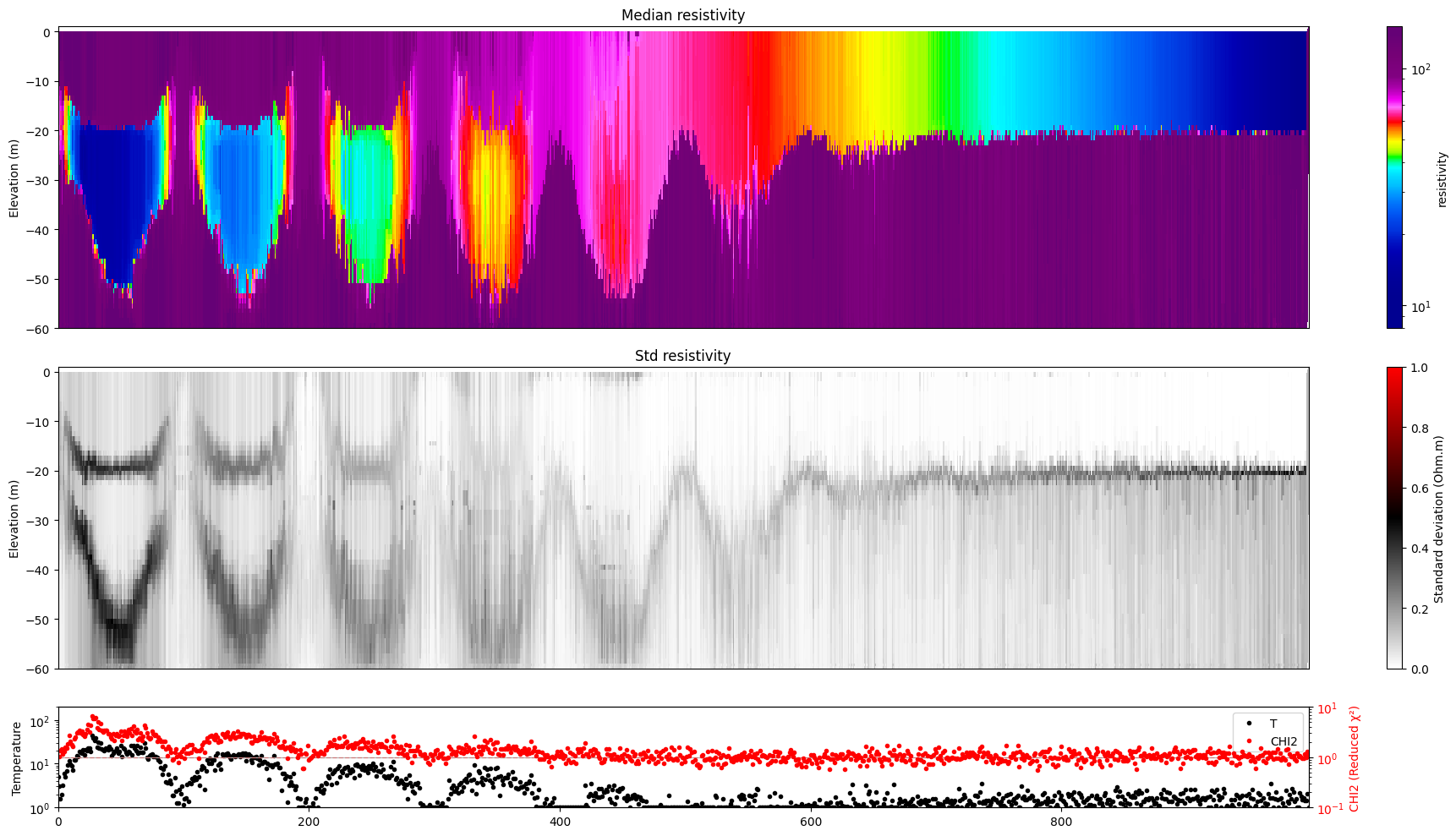
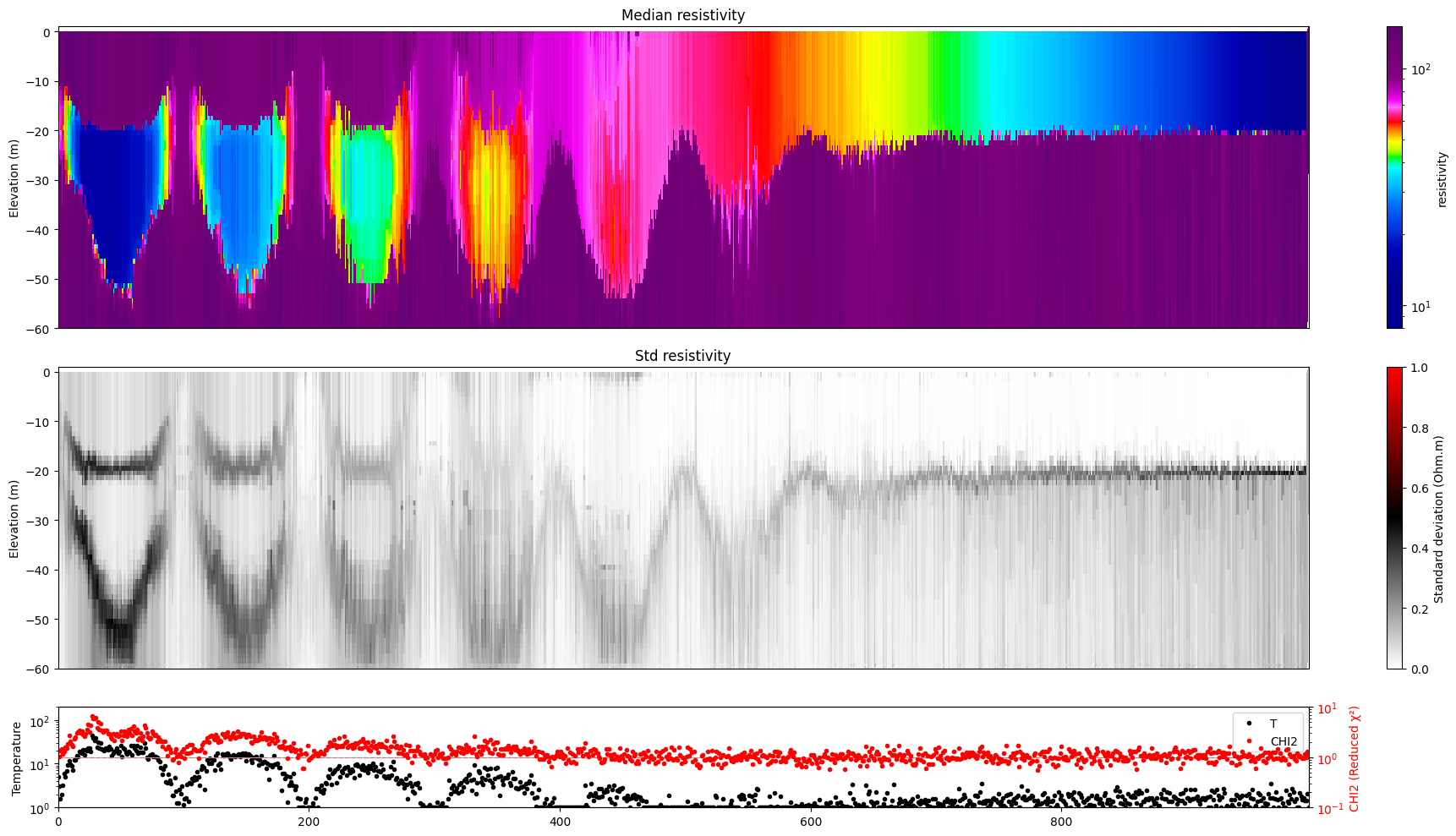
[11]:
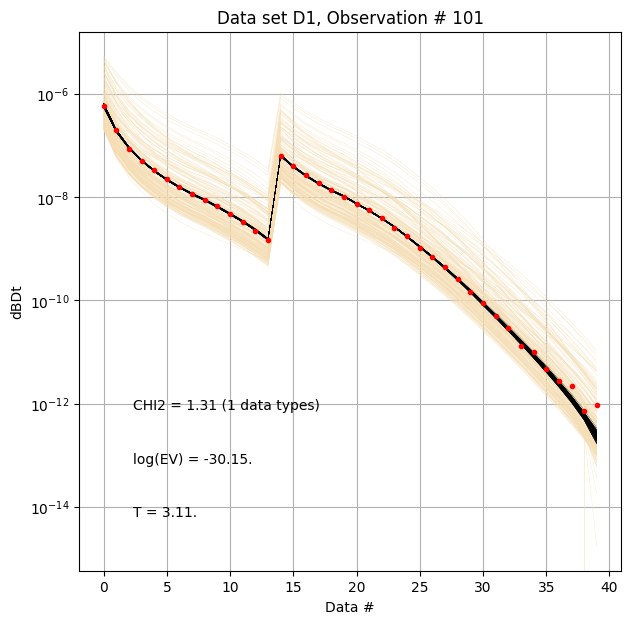
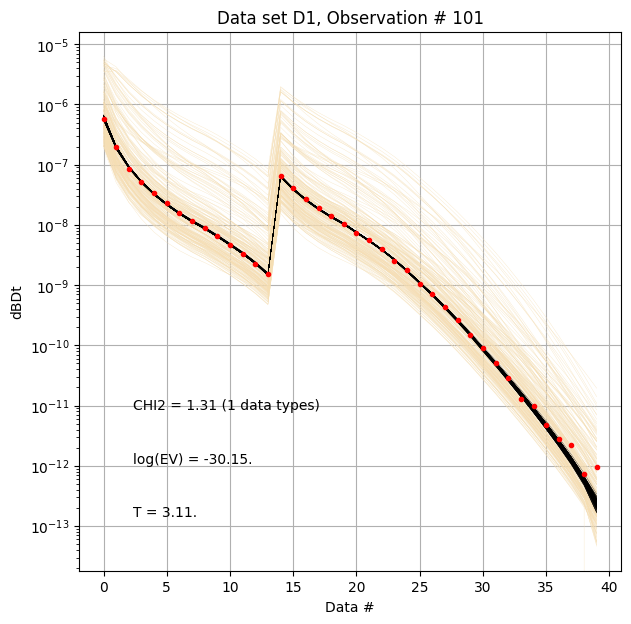
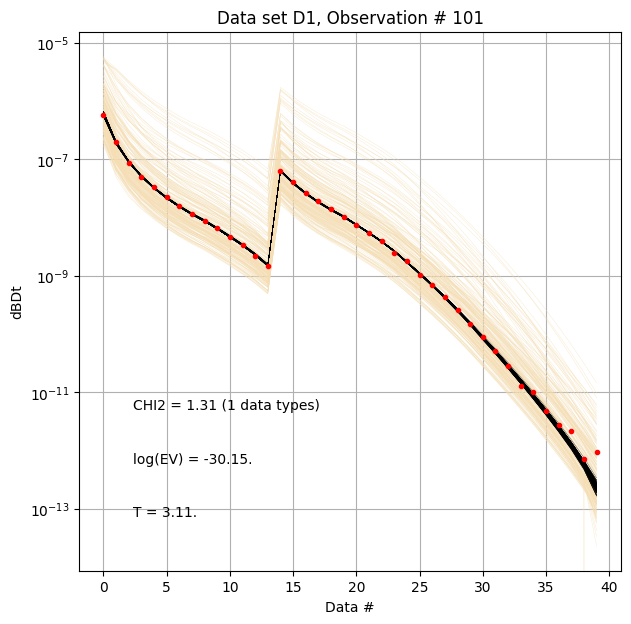
for i in range(len(f_post_h5_arr)):
ig.plot_data_prior_post(f_post_h5_arr[0], i_plot=100, hardcopy=hardcopy)
[12]:
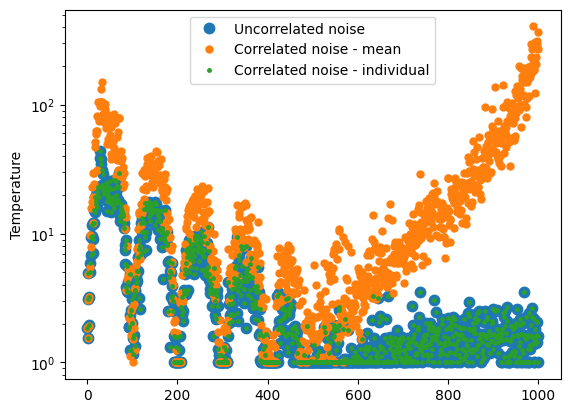
plt.figure()
for i in range(len(T_arr)):
plt.semilogy(T_arr[i], '.', label=name_arr[i], markersize=15-5*i)
plt.legend()
plt.ylabel('Temperature')
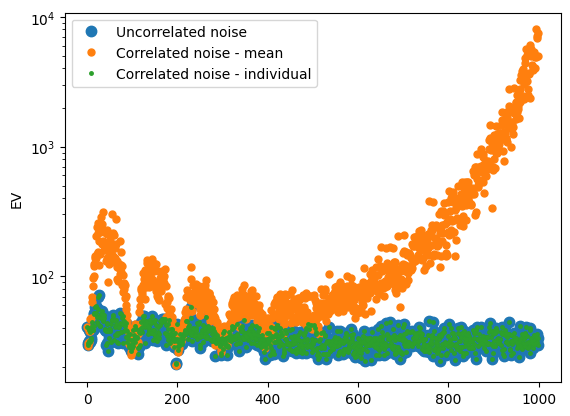
plt.figure()
for i in range(len(T_arr)):
plt.semilogy(-EV_arr[i], '.', label=name_arr[i], markersize=15-5*i)
plt.legend()
plt.ylabel('EV')
plt.figure()
for i in range(len(T_arr)):
plt.semilogy(EV_post_arr[i], '.', label=name_arr[i], markersize=15-5*i)
plt.legend()
plt.ylabel('EV_post')
[12]:
Text(0, 0.5, 'EV_post')
Data in the log-space, and correlated Gaussian noise¶
The data can be transformed to the log-space, and the noise model can be applied in the log-space.
[13]:
# Add constant covariance to Cd -->
# A simple way to introduce correlated noise
corrlev = 0.02**2
lD_obs = np.log10(D_ref)
lD_std_up = np.abs(np.log10(D_ref+D_std)-lD_obs)
lD_std_down = np.abs(np.log10(D_ref-D_std)-lD_obs)
lD_std = np.abs((lD_std_up+lD_std_down)/2) + np.sqrt(corrlev)
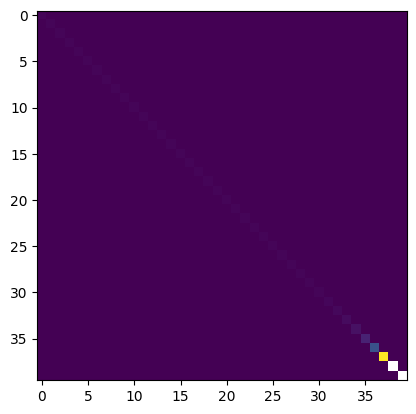
lCd_single = np.diag(np.mean(lD_std, axis=0)**2)+corrlev
plt.imshow(lCd_single)
ns,nd=D_std.shape
lCd_mul = np.zeros((ns,nd,nd))
for i in range(ns):
lCd_mul[i] = np.diag(lD_std[i]**2)+corrlev
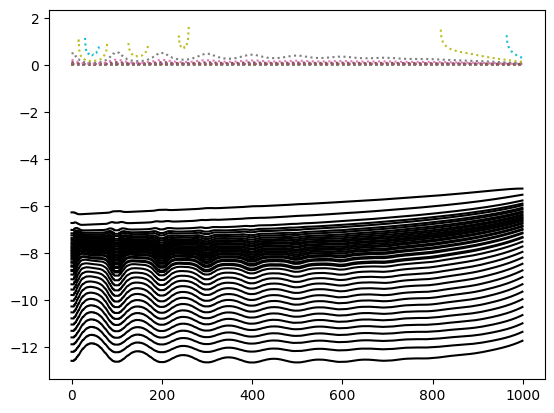
plt.figure()
plt.plot(lD_obs,'k-')
plt.plot(lD_std,':')
f_data_log_1_h5_f_out = ig.save_data_gaussian(lD_obs, D_std = lD_std, f_data_h5 = 'data_log_uncorr', id=1, showInfo=0, is_log=1, delete_if_exist=True)
f_data_log_2_h5_f_out = ig.save_data_gaussian(lD_obs, Cd = lCd_single, f_data_h5 = 'data_log_corr', id=1, showInfo=0, is_log=1, delete_if_exist=True)
f_data_log_3_h5_f_out = ig.save_data_gaussian(lD_obs, Cd = lCd_mul, f_data_h5 = 'data_log_corr2', id=1, showInfo=0, is_log=1, delete_if_exist=True)
f_data_arr = [f_data_log_1_h5_f_out,f_data_log_2_h5_f_out,f_data_log_3_h5_f_out]
Data has 1000 stations and 40 channels
Adding group data_log_uncorr:D1
Data has 1000 stations and 40 channels
Adding group data_log_corr:D1
Data has 1000 stations and 40 channels
Adding group data_log_corr2:D1
[14]:
lCd_single
[14]:
array([[0.00149119, 0.0004 , 0.0004 , ..., 0.0004 , 0.0004 ,
0.0004 ],
[0.0004 , 0.00149124, 0.0004 , ..., 0.0004 , 0.0004 ,
0.0004 ],
[0.0004 , 0.0004 , 0.00149133, ..., 0.0004 , 0.0004 ,
0.0004 ],
...,
[0.0004 , 0.0004 , 0.0004 , ..., 0.08107004, 0.0004 ,
0.0004 ],
[0.0004 , 0.0004 , 0.0004 , ..., 0.0004 , nan,
0.0004 ],
[0.0004 , 0.0004 , 0.0004 , ..., 0.0004 , 0.0004 ,
nan]], shape=(40, 40))
[15]:
recomputePriorData = False
if recomputePriorData:
f_prior_log_data_h5 = ig.prior_data_gaaem(f_prior_h5, file_gex, N=N-1, is_log=True)
else:
# Simple load the old data and save it in log-space
f_prior_log_data_h5 = 'd_log.h5'
ig.copy_hdf5_file(f_prior_h5, f_prior_log_data_h5)
D, idx = ig.load_prior_data(f_prior_data_h5);
Dlog = np.log10(D[0])
ig.save_prior_data(f_prior_log_data_h5, Dlog, id=1)
Loading prior data from PRIOR_UNIFORM_NL_3-3_uniform_N2000000_TX07_20231016_2x4_RC20-33_Nh280_Nf12.h5. Using prior data ids: []
- /D1: N,nd = 2000000/40
[16]:
f_post_log_h5_arr = []
for i in range(len(f_data_arr)):
f_data_h5 = f_data_arr[i]
f_post_h5 = ig.integrate_rejection(f_prior_log_data_h5, f_data_h5,
parallel=parallel,
Ncpu=8,
nr=1000,
updatePostStat = True,
)
f_post_log_h5_arr.append(f_post_h5)
integrate_rejection: Time=299.6s/1000 soundings, 299.6ms/sounding, 3.3it/s. T_av=1.3, EV_av=-11.5
integrate_rejection: Time=381.9s/1000 soundings, 381.9ms/sounding, 2.6it/s. T_av=1.1, EV_av=-10.1
integrate_rejection: Time=394.7s/1000 soundings, 394.7ms/sounding, 2.5it/s. T_av=1.1, EV_av=-10.2
[17]:
for i in range(len(f_post_log_h5_arr)):
ig.plot_profile(f_post_log_h5_arr[i],hardcopy=hardcopy, clim = clim, im=1)
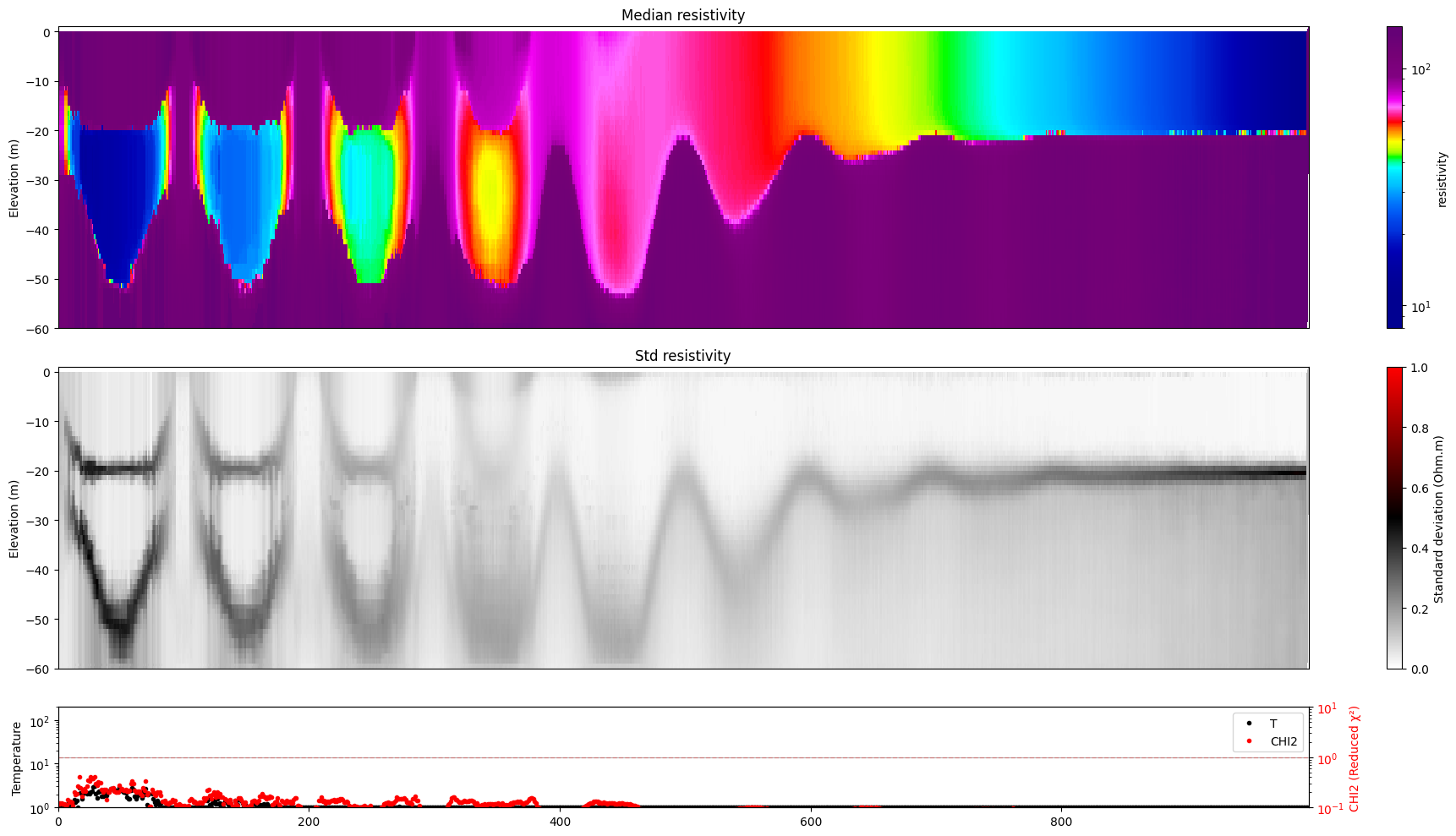
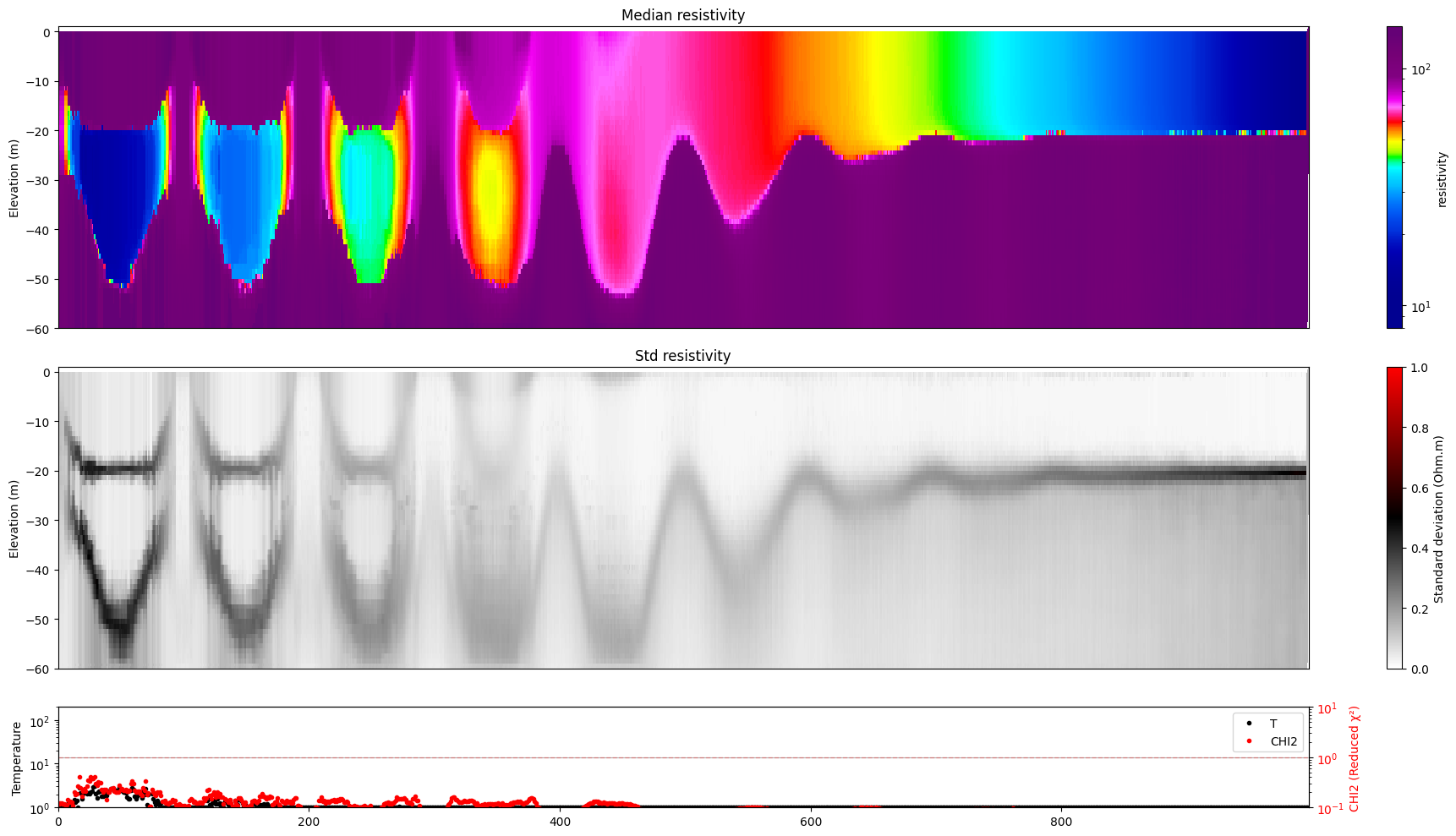
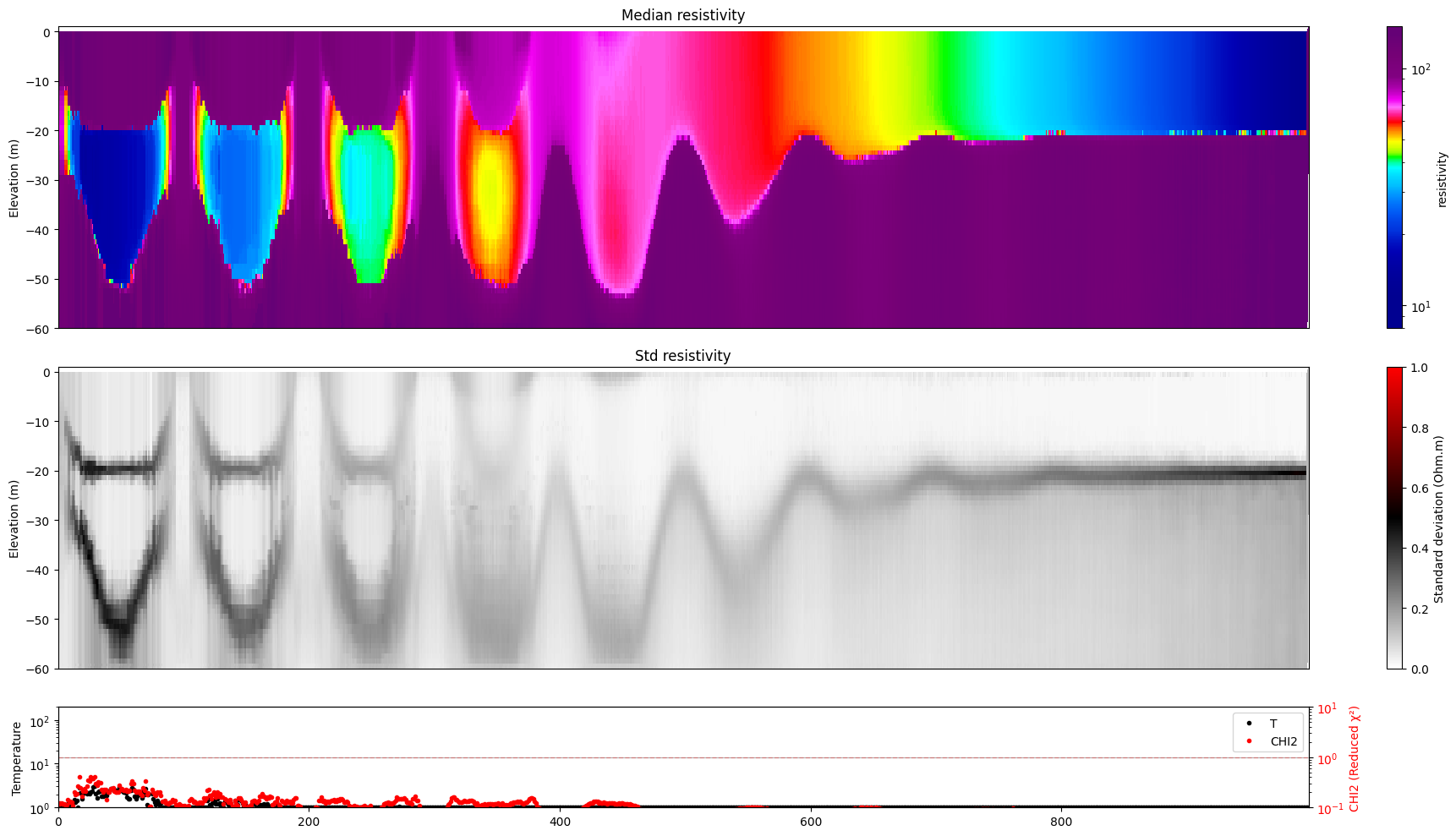
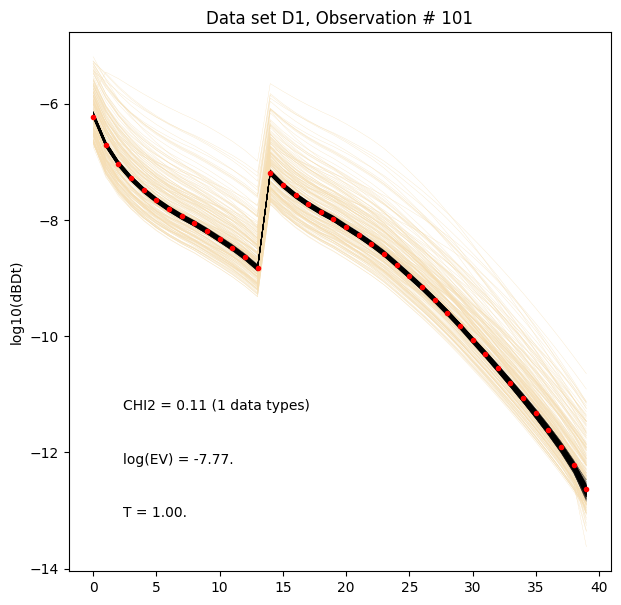
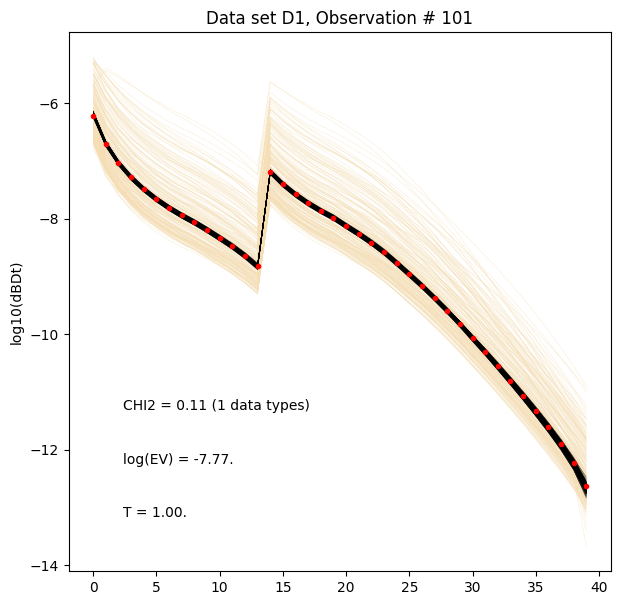
[18]:
for i in range(len(f_post_log_h5_arr)):
ig.plot_data_prior_post(f_post_log_h5_arr[i], i_plot=100, hardcopy=hardcopy, is_log=True)
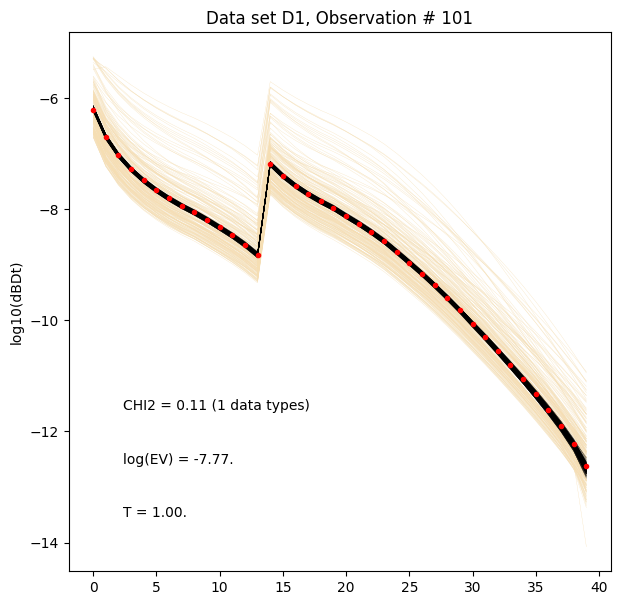
[ ]:
[ ]: